RadD is a RecA-dependent accessory protein that accelerates DNA strand exchange
- PMID: 35150260
- PMCID: PMC8887467
- DOI: 10.1093/nar/gkac041
RadD is a RecA-dependent accessory protein that accelerates DNA strand exchange
Abstract
In rapidly growing cells, with recombinational DNA repair required often and a new replication fork passing every 20 min, the pace of RecA-mediated DNA strand exchange is potentially much too slow for bacterial DNA metabolism. The enigmatic RadD protein, a putative SF2 family helicase, exhibits no independent helicase activity on branched DNAs. Instead, RadD greatly accelerates RecA-mediated DNA strand exchange, functioning only when RecA protein is present. The RadD reaction requires the RadD ATPase activity, does not require an interaction with SSB, and may disassemble RecA filaments as it functions. We present RadD as a new class of enzyme, an accessory protein that accelerates DNA strand exchange, possibly with a helicase-like action, in a reaction that is entirely RecA-dependent. RadD is thus a DNA strand exchange (recombination) synergist whose primary function is to coordinate closely with and accelerate the DNA strand exchange reactions promoted by the RecA recombinase. Multiple observations indicate a uniquely close coordination of RadD with RecA function.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
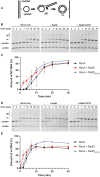
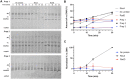

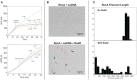
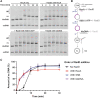
Similar articles
-
Interaction with single-stranded DNA-binding protein modulates Escherichia coli RadD DNA repair activities.J Biol Chem. 2023 Jun;299(6):104773. doi: 10.1016/j.jbc.2023.104773. Epub 2023 May 2. J Biol Chem. 2023. PMID: 37142225 Free PMC article.
-
Quantitative RecA protein binding to the hybrid duplex product of DNA strand exchange.Biochemistry. 1995 Aug 29;34(34):10859-66. doi: 10.1021/bi00034a019. Biochemistry. 1995. PMID: 7662666
-
The Escherichia coli DinD protein modulates RecA activity by inhibiting postsynaptic RecA filaments.J Biol Chem. 2011 Aug 26;286(34):29480-91. doi: 10.1074/jbc.M111.245373. Epub 2011 Jun 22. J Biol Chem. 2011. PMID: 21697094 Free PMC article.
-
The RecA protein as a recombinational repair system.Mol Microbiol. 1991 Jun;5(6):1295-9. doi: 10.1111/j.1365-2958.1991.tb00775.x. Mol Microbiol. 1991. PMID: 1787786 Review.
-
The bacterial RecA protein as a motor protein.Annu Rev Microbiol. 2003;57:551-77. doi: 10.1146/annurev.micro.57.030502.090953. Annu Rev Microbiol. 2003. PMID: 14527291 Review.
Cited by
-
Identification of recG genetic interactions in Escherichia coli by transposon sequencing.J Bacteriol. 2023 Dec 19;205(12):e0018423. doi: 10.1128/jb.00184-23. Epub 2023 Nov 29. J Bacteriol. 2023. PMID: 38019006 Free PMC article.
-
Generation and Repair of Postreplication Gaps in Escherichia coli.Microbiol Mol Biol Rev. 2023 Jun 28;87(2):e0007822. doi: 10.1128/mmbr.00078-22. Epub 2023 May 22. Microbiol Mol Biol Rev. 2023. PMID: 37212693 Free PMC article. Review.
-
Biochemical and Structural Analyses Shed Light on the Mechanisms of RadD DNA Binding and Its ATPase from Escherichia coli.Int J Mol Sci. 2023 Jan 1;24(1):741. doi: 10.3390/ijms24010741. Int J Mol Sci. 2023. PMID: 36614183 Free PMC article.
-
X-ray crystal structure of the Escherichia coli RadD DNA repair protein bound to ADP reveals a novel zinc ribbon domain.PLoS One. 2022 Apr 28;17(4):e0266031. doi: 10.1371/journal.pone.0266031. eCollection 2022. PLoS One. 2022. PMID: 35482735 Free PMC article.
-
Molecular insights into the prototypical single-stranded DNA-binding protein from E. coli.Crit Rev Biochem Mol Biol. 2024 Feb-Apr;59(1-2):99-127. doi: 10.1080/10409238.2024.2330372. Epub 2024 May 21. Crit Rev Biochem Mol Biol. 2024. PMID: 38770626 Free PMC article. Review.
References
-
- Courcelle J., Belle J.J., Courcelle C.T.. When replication travels on damaged templates: bumps and blocks in the road. Res. Microbiol. 2004; 155:231–237. - PubMed
-
- Cox M.M., Goodman M.F., Kreuzer K.N., Sherratt D.J., Sandler S.J., Marians K.J.. The importance of repairing stalled replication forks. Nature. 2000; 404:37–41. - PubMed
-
- Kowalczykowski S.C. Initiation of genetic recombination and recombination-dependent replication. Trends Biochem. Sci. 2000; 25:156–165. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

