Evaluation of process limit of detection and quantification variation of SARS-CoV-2 RT-qPCR and RT-dPCR assays for wastewater surveillance
- PMID: 35152136
- PMCID: PMC8812148
- DOI: 10.1016/j.watres.2022.118132
Evaluation of process limit of detection and quantification variation of SARS-CoV-2 RT-qPCR and RT-dPCR assays for wastewater surveillance
Abstract
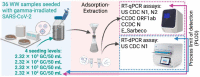
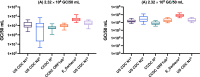
Effective wastewater surveillance of SARS-CoV-2 RNA requires the rigorous characterization of the limit of detection resulting from the entire sampling process - the process limit of detection (PLOD). Yet to date, no studies have gone beyond quantifying the assay limit of detection (ALOD) for RT-qPCR or RT-dPCR assays. While the ALOD is the lowest number of gene copies (GC) associated with a 95% probability of detection in a single PCR reaction, the PLOD represents the sensitivity of the method after considering the efficiency of all processing steps (e.g., sample handling, concentration, nucleic acid extraction, and PCR assays) to determine the number of GC in the wastewater sample matrix with a specific probability of detection. The primary objective of this study was to estimate the PLOD resulting from the combination of primary concentration and extraction with six SARS-CoV-2 assays: five RT-qPCR assays (US CDC N1 and N2, China CDC N and ORF1ab (CCDC N and CCDC ORF1ab), and E_Sarbeco RT-qPCR, and one RT-dPCR assay (US CDC N1 RT-dPCR) using two models (exponential survival and cumulative Gaussian). An adsorption extraction (AE) concentration method (i.e., virus adsorption on membrane and the RNA extraction from the membrane) was used to concentrate gamma-irradiated SARS-CoV-2 seeded into 36 wastewater samples. Overall, the US CDC N1 RT-dPCR and RT-qPCR assays had the lowest ALODs (< 10 GC/reaction) and PLODs (<3,954 GC/50 mL; 95% probability of detection) regardless of the seeding level and model used. Nevertheless, consistent amplification and detection rates decreased when seeding levels were < 2.32 × 103 GC/50 mL even for US CDC N1 RT-qPCR and RT-dPCR assays. Consequently, when SARS-CoV-2 RNA concentrations are expected to be low, it may be necessary to improve the positive detection rates of wastewater surveillance by analyzing additional field and RT-PCR replicates. To the best of our knowledge, this is the first study to assess the SARS-CoV-2 PLOD for wastewater and provides important insights on the analytical limitations for trace detection of SARS-CoV-2 RNA in wastewater.
Keywords: COVID-19; Concentration method; Detection limit; Enveloped virus; Recovery; SARS-CoV-2; Wastewater.
Crown Copyright © 2022. Published by Elsevier Ltd. All rights reserved.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

References
-
- Ahmed W., Hamilton K.A., Lobos A., Hughes B., Staley C., Sadowsky M.J., Harwood V.J. Quantitative microbial risk assessment of microbial source tracking markers in recreational water contaminated with fresh untreated and secondary treated sewage. Environ. Int. 2018;117:243–249. - PubMed
-
- Ahmed W., Angel N., Edson J., Bibby K., Bivins A., Brien J.W.O., Choi P.M., Kitajima M., Simpson S.L., Li J., Tscharke B., Verhagen R., Smith W.J.M., Zaugg J., Dierens L., Hugenholtz P., Thomas K.V., Mueller J.F. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020;728 - PMC - PubMed
-
- Ahmed W., Bivins A., Bertsch P.M., Bibby K., Choi P.M., Farkas K., Gyawali P., Hamilton K.A., Haramoto E., Kitajima M. Surveillance of SARS-CoV-2 RNA in wastewater: Methods optimisation and quality control are crucial for generating reliable public health information. Curr. Opin. Environ. Sci. Health. 2020;17:82–93. - PMC - PubMed
-
- Ahmed W., Bertsch P.M., Angel N., Bibby K., Bivins A., Dierens L., Edson J., Ehret j., Gyawali P., Hamilton K.A., Hosegood I., Hugenholtz P., Jiang G., Kitajima M., Sichani H.T., Shi J., Shimko K.M., Simpson S.L., Smith W.J.M., Symonds E.M., Thomas K.V., Verhagen R., Zaugg J., Mueller J.F. Detection of SARS-CoV-2 RNA in commercial passenger aircraft and cruise ship wastewater: a surveillance tool for assessing the presence of COVID-19 infected travellers. J. Travel Med. 2020;27(5):taaa116. - PMC - PubMed
-
- Ahmed W., Bertsch P.M., Bivins A., Bibby K., Farkas K., Gathercole A., Haramoto E., Gyawali P., Korajkic A., McMinn B.R., Mueller J.F., Simpson S.L., Smith W.J.M., Symonds E.M., Thomas K.V., Verhagen R., Kitajima M. Comparison of virus concentration methods for the RT-qPCR-based recovery of murine hepatitis virus, a surrogate for SARS-CoV-2 from untreated wastewater. Sci. Total Environ. 2020;739 - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

