Cryo-Treatment Enhances the Embryogenicity of Mature Somatic Embryos via the lncRNA-miRNA-mRNA Network in White Spruce
- PMID: 35163033
- PMCID: PMC8834816
- DOI: 10.3390/ijms23031111
Cryo-Treatment Enhances the Embryogenicity of Mature Somatic Embryos via the lncRNA-miRNA-mRNA Network in White Spruce
Abstract
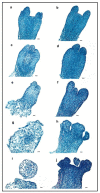
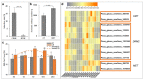
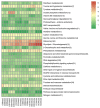
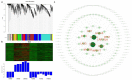
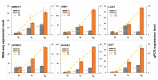
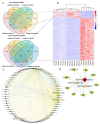
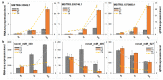
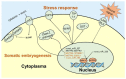
In conifers, somatic embryogenesis is uniquely initiated from immature embryos in a narrow time window, which is considerably hindered by the difficulty to induce embryogenic tissue (ET) from other tissues, including mature somatic embryos. In this study, the embryogenic ability of newly induced ET and DNA methylation levels was detected, and whole-transcriptome sequencing analyses were carried out. The results showed that ultra-low temperature treatment significantly enhanced ET induction from mature somatic embryos, with the induction rate from 0.4% to 15.5%, but the underlying mechanisms remain unclear. The newly induced ET showed higher capability in generating mature embryos than the original ET. DNA methylation levels fluctuated during the ET induction process. Here, WGCNA analysis revealed that OPT4, TIP1-1, Chi I, GASA5, GST, LAX3, WRKY7, MYBS3, LRR-RLK, PBL7, and WIN1 genes are involved in stress response and auxin signal transduction. Through co-expression analysis, lncRNAs MSTRG.505746.1, MSTRG.1070680.1, and MSTRG.33602.1 might bind to pre-novel_miR_339 to promote the expression of WRKY7 genes for stress response; LAX3 could be protected by lncRNAs MSTRG.1070680.1 and MSTRG.33602.1 via serving as sponges for novel_miR_495 to initiate auxin signal transduction; lncRNAs MSTRG.505746.1, MSTRG.1070680.1, and MSTRG.33602.1 might serve as sponges for novel_miR_527 to enhance the expression of Chi I for early somatic embryo development. This study provides new insight into the area of stress-enhanced early somatic embryogenesis in conifers, which is also attributable to practical applications.
Keywords: conifer; cryo-treated mature somatic embryos; liquid nitrogen; lncRNAs; miRNAs; whole transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Attree S.M., Fowke L.C. Embryogeny of gymnosperms—Advances in synthetic seed technology of conifers. Plant Cell Tissue Organ. Cult. 1993;35:31–35. doi: 10.1007/BF00043936. - DOI
-
- Kong L., von Aderkas P. A novel method of cryopreservation without a cryoprotectant for immature somatic embryos of conifer. Plant. Cell Tissue Organ. Cult. 2010;106:115–125. doi: 10.1007/s11240-010-9899-x. - DOI
-
- Tremblay F.M. Somatic embryogenesis and plantlet regeneration from embryos isolated from stored seeds of Picea glauca. Can. J. Bot. 1990;68:236–242. doi: 10.1139/b90-033. - DOI
MeSH terms
Substances
Grants and funding
- 2017YDF0600404-1/National Key R&D Program of China
- Beijing Advanced Innovation Center for Tree Breeding by Molecular Design/Somatic embryogenesis and high efficient propagation technology in trees
- 31901289/the National Natural Science Foundation of China
- 20326333D/Key R&D Program of Heibei Province, China
- 20170112006/Major Science and Technology Special Project of Xuchang, Henan province, China
LinkOut - more resources
Full Text Sources
Research Materials

