Induction of the Unfolded Protein Response at High Temperature in Saccharomyces cerevisiae
- PMID: 35163590
- PMCID: PMC8836091
- DOI: 10.3390/ijms23031669
Induction of the Unfolded Protein Response at High Temperature in Saccharomyces cerevisiae
Abstract
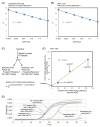
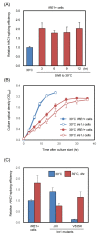
Ire1 is an endoplasmic reticulum (ER)-located endoribonuclease that is activated in response to ER stress. In yeast Saccharomyces cerevisiae cells, Ire1 promotes HAC1-mRNA splicing to remove the intron sequence from the HAC1u mRNA ("u" stands for "uninduced"). The resulting mRNA, which is named HAC1i mRNA ("i" stands for "induced"), is then translated into a transcription factor that is involved in the unfolded protein response (UPR). In this study, we designed an oligonucleotide primer that specifically hybridizes to the exon-joint site of the HAC1i cDNA. This primer allowed us to perform real-time reverse transcription-PCR to quantify HAC1i mRNA abundance with high sensitivity. Using this method, we detected a minor induction of HAC1-mRNA splicing in yeast cells cultured at their maximum growth temperature of 39 °C. Based on our analyses of IRE1-gene mutant strains, we propose that when yeast cells are cultured at or near their maximum growth temperature, protein folding in the ER is disturbed, leading to a minor UPR induction that supports cellular growth.
Keywords: PCR; endoplasmic reticulum; mRNA solicing; yeast.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Fast-Growing Saccharomyces cerevisiae Cells with a Constitutive Unfolded Protein Response and Their Potential for Lipidic Molecule Production.Appl Environ Microbiol. 2022 Nov 8;88(21):e0108322. doi: 10.1128/aem.01083-22. Epub 2022 Oct 18. Appl Environ Microbiol. 2022. PMID: 36255243 Free PMC article.
-
Ribosome depurination by ricin leads to inhibition of endoplasmic reticulum stress-induced HAC1 mRNA splicing on the ribosome.J Biol Chem. 2019 Nov 22;294(47):17848-17862. doi: 10.1074/jbc.RA119.009128. Epub 2019 Oct 17. J Biol Chem. 2019. PMID: 31624149 Free PMC article.
-
IRE1- and HAC1-independent transcriptional regulation in the unfolded protein response of yeast.Mol Microbiol. 2003 Aug;49(3):591-606. doi: 10.1046/j.1365-2958.2003.03585.x. Mol Microbiol. 2003. PMID: 12864846
-
Translation Control of HAC1 by Regulation of Splicing in Saccharomyces cerevisiae.Int J Mol Sci. 2019 Jun 12;20(12):2860. doi: 10.3390/ijms20122860. Int J Mol Sci. 2019. PMID: 31212749 Free PMC article. Review.
-
The unfolded protein response: the dawn of a new field.Proc Jpn Acad Ser B Phys Biol Sci. 2015;91(9):469-80. doi: 10.2183/pjab.91.469. Proc Jpn Acad Ser B Phys Biol Sci. 2015. PMID: 26560836 Free PMC article. Review.
Cited by
-
Transcriptome-wide mRNA condensation precedes stress granule formation and excludes stress-induced transcripts.bioRxiv [Preprint]. 2024 May 2:2024.04.15.589678. doi: 10.1101/2024.04.15.589678. bioRxiv. 2024. PMID: 38659805 Free PMC article. Preprint.
-
Induction of Endoplasmic Reticulum Stress by Prodigiosin in Yeast Saccharomyces cerevisiae.Curr Issues Mol Biol. 2024 Feb 26;46(3):1768-1776. doi: 10.3390/cimb46030116. Curr Issues Mol Biol. 2024. PMID: 38534732 Free PMC article.
-
Fast-Growing Saccharomyces cerevisiae Cells with a Constitutive Unfolded Protein Response and Their Potential for Lipidic Molecule Production.Appl Environ Microbiol. 2022 Nov 8;88(21):e0108322. doi: 10.1128/aem.01083-22. Epub 2022 Oct 18. Appl Environ Microbiol. 2022. PMID: 36255243 Free PMC article.
-
Endoplasmic stress sensor Ire1 is involved in cytosolic/nuclear protein quality control in Pichia pastoris cells independent of HAC1.Front Microbiol. 2023 Jun 20;14:1157146. doi: 10.3389/fmicb.2023.1157146. eCollection 2023. Front Microbiol. 2023. PMID: 37415818 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

