The Absence of Retroelement Activity Is Characteristic for Childhood Acute Leukemias and Adult Acute Lymphoblastic Leukemia
- PMID: 35163677
- PMCID: PMC8835895
- DOI: 10.3390/ijms23031756
The Absence of Retroelement Activity Is Characteristic for Childhood Acute Leukemias and Adult Acute Lymphoblastic Leukemia
Abstract
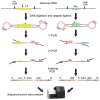
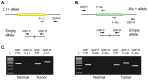
Retroelements (RE) have been proposed as important players in cancerogenesis. Different cancer types are characterized by a different level of tumor-specific RE insertions. In previous studies, small cohorts of hematological malignancies, such as acute myeloid leukemia, multiple myeloma, and chronic lymphocytic leukemia have been characterized by a low level of RE insertional activity. Acute lymphoblastic leukemia (ALL) in adults and childhood acute leukemias have not been studied in this context. We performed a search for new RE insertions (Alu and L1) in 44 childhood ALL, 14 childhood acute myeloid leukemia, and 14 adult ALL samples using a highly sensitive NGS-based approach. First, we evaluated the method sensitivity revealing the 1% detection threshold for the proportion of cells with specific RE insertion. Following this result, we did not identify new tumor-specific RE insertions in the tested cohort of acute leukemia samples at the established level of sensitivity. Additionally, we analyzed the transcription levels of active L1 copies and found them increased. Thus, the increased transcription of active L1 copies is not sufficient for overt elevation of L1 retrotranspositional activity in leukemia.
Keywords: acute leukemia; mobile elements; retroelements; tumor-specific insertions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Comparison of minimal residual disease (MRD) monitoring by WT1 quantification between childhood acute myeloid leukemia and acute lymphoblastic leukemia.Eur Rev Med Pharmacol Sci. 2015;19(14):2679-88. Eur Rev Med Pharmacol Sci. 2015. PMID: 26221900
-
[The functional analysis of polymorphic insertions of Alu retroelements at acute lymphoblastic leukemia].Bioorg Khim. 2012 May-Jun;38(3):351-64. doi: 10.1134/s1068162012030089. Bioorg Khim. 2012. PMID: 22997707 Russian.
-
mRNA expression of the XAGE-1 gene in human acute leukemia.Int J Hematol. 2010 Mar;91(2):209-12. doi: 10.1007/s12185-010-0527-7. Epub 2010 Feb 24. Int J Hematol. 2010. PMID: 20178013
-
Childhood and adolescent lymphoid and myeloid leukemia.Hematology Am Soc Hematol Educ Program. 2004:118-45. doi: 10.1182/asheducation-2004.1.118. Hematology Am Soc Hematol Educ Program. 2004. PMID: 15561680 Review.
-
Clinical cytogenetics in pediatric acute leukemia: an update.Clin Lymphoma Myeloma Leuk. 2012 Aug;12(4):230-7. doi: 10.1016/j.clml.2012.04.004. Epub 2012 May 19. Clin Lymphoma Myeloma Leuk. 2012. PMID: 22609262 Review.
Cited by
-
A Glance at Molecular Advances in Cancer Genetics: A Baffling Puzzle Still to Be Solved.Int J Mol Sci. 2023 Jan 11;24(2):1394. doi: 10.3390/ijms24021394. Int J Mol Sci. 2023. PMID: 36674909 Free PMC article.
References
-
- Lander E.S., Linton L.M., Birren B., Nusbaum C., Zody M.C., Baldwin J., Devon K., Dewar K., Doyle M., FitzHugh W., et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

