Structural and Biochemical Analysis of the Furan Aldehyde Reductase YugJ from Bacillus subtilis
- PMID: 35163804
- PMCID: PMC8836905
- DOI: 10.3390/ijms23031882
Structural and Biochemical Analysis of the Furan Aldehyde Reductase YugJ from Bacillus subtilis
Abstract
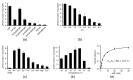
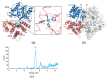
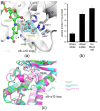
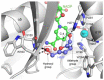
NAD(H)/NADP(H)-dependent aldehyde/alcohol oxidoreductase (AAOR) participates in a wide range of physiologically important cellular processes by reducing aldehydes or oxidizing alcohols. Among AAOR substrates, furan aldehyde is highly toxic to microorganisms. To counteract the toxic effect of furan aldehyde, some bacteria have evolved AAOR that converts furan aldehyde into a less toxic alcohol. Based on biochemical and structural analyses, we identified Bacillus subtilis YugJ as an atypical AAOR that reduces furan aldehyde. YugJ displayed high substrate specificity toward 5-hydroxymethylfurfural (HMF), a furan aldehyde, in an NADPH- and Ni2+-dependent manner. YugJ folds into a two-domain structure consisting of a Rossmann-like domain and an α-helical domain. YugJ interacts with NADP and Ni2+ using the interdomain cleft of YugJ. A comparative analysis of three YugJ structures indicated that NADP(H) binding plays a key role in modulating the interdomain dynamics of YugJ. Noticeably, a nitrate ion was found in proximity to the nicotinamide ring of NADP in the YugJ structure, and the HMF-reducing activity of YugJ was inhibited by nitrate, providing insights into the substrate-binding mode of YugJ. These findings contribute to the characterization of the YugJ-mediated furan aldehyde reduction mechanism and to the rational design of improved furan aldehyde reductases for the biofuel industry.
Keywords: 5-hydroxymethylfurfural; Bacillus subtilis; NADPH cofactor; Ni2+ cofactor; YugJ; crystal structure; furan aldehyde reductase.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Kinetic mechanism of an aldehyde reductase of Saccharomyces cerevisiae that relieves toxicity of furfural and 5-hydroxymethylfurfural.Biochim Biophys Acta. 2011 Dec;1814(12):1686-94. doi: 10.1016/j.bbapap.2011.08.011. Epub 2011 Aug 26. Biochim Biophys Acta. 2011. PMID: 21890004
-
A novel NADPH-dependent aldehyde reductase gene from Saccharomyces cerevisiae NRRL Y-12632 involved in the detoxification of aldehyde inhibitors derived from lignocellulosic biomass conversion.Gene. 2009 Oct 1;446(1):1-10. doi: 10.1016/j.gene.2009.06.018. Epub 2009 Jul 3. Gene. 2009. PMID: 19577617
-
YNL134C from Saccharomyces cerevisiae encodes a novel protein with aldehyde reductase activity for detoxification of furfural derived from lignocellulosic biomass.Yeast. 2015 May;32(5):409-22. doi: 10.1002/yea.3068. Epub 2015 Mar 3. Yeast. 2015. PMID: 25656244
-
Functions of aldehyde reductases from Saccharomyces cerevisiae in detoxification of aldehyde inhibitors and their biotechnological applications.Appl Microbiol Biotechnol. 2018 Dec;102(24):10439-10456. doi: 10.1007/s00253-018-9425-3. Epub 2018 Oct 10. Appl Microbiol Biotechnol. 2018. PMID: 30306200 Review.
-
Aldose reductase: monosaccharide autoxidation and NADPH binding.Biochem Soc Trans. 1996 Aug;24(3):888-91. doi: 10.1042/bst0240888. Biochem Soc Trans. 1996. PMID: 8878869 Review. No abstract available.
Cited by
-
A model industrial workhorse: Bacillus subtilis strain 168 and its genome after a quarter of a century.Microb Biotechnol. 2023 Jun;16(6):1203-1231. doi: 10.1111/1751-7915.14257. Epub 2023 Apr 1. Microb Biotechnol. 2023. PMID: 37002859 Free PMC article. Review.
-
Exploring the Potential of Phytocannabinoids Against Multidrug-Resistant Bacteria.Plants (Basel). 2025 Jun 20;14(13):1901. doi: 10.3390/plants14131901. Plants (Basel). 2025. PMID: 40647911 Free PMC article.
-
Escherichia coli alcohol dehydrogenase YahK is a protein that binds both iron and zinc.PeerJ. 2024 Sep 10;12:e18040. doi: 10.7717/peerj.18040. eCollection 2024. PeerJ. 2024. PMID: 39282118 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

