Genome-Wide Characterization of SARS-CoV-2 Cytopathogenic Proteins in the Search of Antiviral Targets
- PMID: 35164548
- PMCID: PMC8844912
- DOI: 10.1128/mbio.00169-22
Genome-Wide Characterization of SARS-CoV-2 Cytopathogenic Proteins in the Search of Antiviral Targets
Abstract
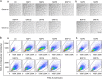
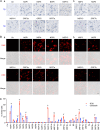
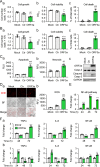
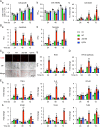
Therapeutic inhibition of critical viral functions is important for curtailing coronavirus disease 2019 (COVID-19). We sought to identify antiviral targets through the genome-wide characterization of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) proteins that are crucial for viral pathogenesis and that cause harmful cytopathogenic effects. All 29 viral proteins were tested in a fission yeast cell-based system using inducible gene expression. Twelve proteins, including eight nonstructural proteins (NSP1, NSP3, NSP4, NSP5, NSP6, NSP13, NSP14, and NSP15) and four accessory proteins (ORF3a, ORF6, ORF7a, and ORF7b), were identified that altered cellular proliferation and integrity and induced cell death. Cell death correlated with the activation of cellular oxidative stress. Of the 12 proteins, ORF3a was chosen for further study in mammalian cells because it plays an important role in viral pathogenesis and its activities are linked to lung tissue damage and a cytokine storm. In human pulmonary and kidney epithelial cells, ORF3a induced cellular oxidative stress associated with apoptosis and necrosis and caused activation of proinflammatory response with production of the cytokines tumor necrosis factor alpha (TNF-α), interleukin-6 (IL-6), and IFN-β1, possibly through the activation of nuclear factor kappa B (NF-κB). To further characterize the mechanism, we tested a natural ORF3a Beta variant, Q57H, and a mutant with deletion of the highly conserved residue, ΔG188. Compared with wild-type ORF3a, the ΔG188 variant yielded more robust activation of cellular oxidative stress, cell death, and innate immune response. Since cellular oxidative stress and inflammation contribute to cell death and tissue damage linked to the severity of COVID-19, our findings suggest that ORF3a is a promising, novel therapeutic target against COVID-19. IMPORTANCE The ongoing COVID-19 pandemic caused by SARS-CoV-2 has claimed over 5.5 million lives with more than 300 million people infected worldwide. While vaccines are effective, the emergence of new viral variants could jeopardize vaccine protection. Treatment of COVID-19 by antiviral drugs provides an alternative to battle against the disease. The goal of this study was to identify viral therapeutic targets that can be used in antiviral drug discovery. Utilizing a genome-wide functional analysis in a fission yeast cell-based system, we identified 12 viral candidates, including ORF3a, which cause cellular oxidative stress, inflammation, apoptosis, and necrosis that contribute to cytopathogenicity and COVID-19. Our findings indicate that antiviral agents targeting ORF3a could have a great impact on COVID-19.
Keywords: ORF3a; SARS-CoV-2; Schizosaccharomyces pombe; apoptosis and necrosis; fission yeast; oxidative stress; proinflammatory response; viral therapeutic target.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Update of
-
Genome-wide characterization of SARS-CoV-2 cytopathogenic proteins in the search of antiviral targets.bioRxiv [Preprint]. 2021 Dec 14:2021.11.23.469747. doi: 10.1101/2021.11.23.469747. bioRxiv. 2021. Update in: mBio. 2022 Feb 15;13(1):e0016922. doi: 10.1128/mbio.00169-22. PMID: 34845452 Free PMC article. Updated. Preprint.
References
-
- Cameroni E, Bowen JE, Rosen LE, Saliba C, Zepeda SK, Culap K, Pinto D, VanBlargan LA, De Marco A, di Iulio J, Zatta F, Kaiser H, Noack J, Farhat N, Czudnochowski N, Havenar-Daughton C, Sprouse KR, Dillen JR, Powell AE, Chen A, Maher C, Yin L, Sun D, Soriaga L, Bassi J, Silacci-Fregni C, Gustafsson C, Franko NM, Logue J, Iqbal NT, Mazzitelli I, Geffner J, Grifantini R, Chu H, Gori A, Riva A, Giannini O, Ceschi A, Ferrari P, Cippà PE, Franzetti-Pellanda A, Garzoni C, Halfmann PJ, Kawaoka Y, Hebner C, Purcell LA, Piccoli L, Pizzuto MS, Walls AC, Diamond MS, et al. 2021. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature doi: 10.1038/s41586-021-04386-2. - DOI - PMC - PubMed
-
- Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O’Meara MJ, Rezelj VV, Guo JZ, Swaney DL, Tummino TA, Hüttenhain R, Kaake RM, Richards AL, Tutuncuoglu B, Foussard H, Batra J, Haas K, Modak M, Kim M, Haas P, Polacco BJ, Braberg H, Fabius JM, Eckhardt M, Soucheray M, Bennett MJ, Cakir M, McGregor MJ, Li Q, Meyer B, Roesch F, Vallet T, Mac Kain A, Miorin L, Moreno E, Naing ZZC, Zhou Y, Peng S, Shi Y, Zhang Z, Shen W, Kirby IT, Melnyk JE, Chorba JS, Lou K, Dai SA, Barrio-Hernandez I, Memon D, Hernandez-Armenta C, et al. 2020. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583:459–468. doi: 10.1038/s41586-020-2286-9. - DOI - PMC - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous
