Soil pH Filters the Association Patterns of Aluminum-Tolerant Microorganisms in Rice Paddies
- PMID: 35166564
- PMCID: PMC8845571
- DOI: 10.1128/msystems.01022-21
Soil pH Filters the Association Patterns of Aluminum-Tolerant Microorganisms in Rice Paddies
Abstract
Soil microbes are considered the second genome of plants. Understanding the distribution and network of aluminum (Al)-tolerant microorganisms is helpful to alleviate Al toxicity to plants in acidic soils. Here, we examined soluble Al3+ and bacterial communities carrying Al resistance genes in paddy soils with a soil pH range of 3.6 to 8.7. In the acidic soil with pH <5.1, the content of Al3+ increased significantly. There were abundant and diverse Al-tolerant microorganisms in acidic soils, including Clostridium, Bacillus, Paenibacillus, Desulfitobacterium, and Desulfosporosinus, etc. Moreover, compared with neutral and alkaline soils, the network structure of Al-tolerant microorganisms was more complex. The potential roles of major Al-tolerant microbial taxa on each other in the ecological network were identified by a directed network along 0.01 pH steps. The influential taxa in the network had a broader niche and contained more antioxidant functional genes to resist Al stress, indicating their survival advantage over the sensitive taxa. Our study is the first to explore the distribution of Al-tolerant microorganisms in continental paddies and reveal their potential associations mediated by pH, which provides a basis for further utilization of microbial resources in acidic agricultural soils. IMPORTANCE Aluminum (Al) toxicity is the primary limiting factor of crop production in acidic soils with pH <5.0. Numerous studies have focused on the mechanism of Al toxicity and tolerance in plants; however, the effects of Al toxicity on soil microorganisms and their tolerance remain less studied. This study investigated the distribution and association patterns of Al-tolerant microorganisms across continental paddy fields with a soil pH range of 3.6 to 8.7. The results showed that soil pH filters exchangeable Al3+ content, diversity, and potential associations of Al-tolerant microbial community. The influential taxa in community network play an important role in Al tolerance and have potential applications in mitigating Al toxicity and promoting crop growth in acidic soils.
Keywords: Al-tolerant bacteria; aluminum toxicity; directed network; functional genes; niche breadth.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


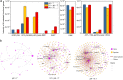


Similar articles
-
Distinct Patterns of Rhizosphere Microbiota Associated With Rice Genotypes Differing in Aluminum Tolerance in an Acid Sulfate Soil.Front Microbiol. 2022 Jun 17;13:933722. doi: 10.3389/fmicb.2022.933722. eCollection 2022. Front Microbiol. 2022. PMID: 35783428 Free PMC article.
-
Toxicity of soil labile aluminum fractions and aluminum species in soil water extracts on the rhizosphere bacterial community of tall fescue.Ecotoxicol Environ Saf. 2020 Jan 15;187:109828. doi: 10.1016/j.ecoenv.2019.109828. Epub 2019 Oct 19. Ecotoxicol Environ Saf. 2020. PMID: 31639644
-
Root proteome of rice studied by iTRAQ provides integrated insight into aluminum stress tolerance mechanisms in plants.J Proteomics. 2014 Feb 26;98:189-205. doi: 10.1016/j.jprot.2013.12.023. Epub 2014 Jan 9. J Proteomics. 2014. PMID: 24412201
-
Toxicity and tolerance of aluminum in plants: tailoring plants to suit to acid soils.Biometals. 2016 Apr;29(2):187-210. doi: 10.1007/s10534-016-9910-z. Epub 2016 Jan 21. Biometals. 2016. PMID: 26796895 Review.
-
Importance of Mineral Nutrition for Mitigating Aluminum Toxicity in Plants on Acidic Soils: Current Status and Opportunities.Int J Mol Sci. 2018 Oct 8;19(10):3073. doi: 10.3390/ijms19103073. Int J Mol Sci. 2018. PMID: 30297682 Free PMC article. Review.
Cited by
-
The Rice-Microbe Nexus: Unlocking Productivity Through Soil Science.Rice (N Y). 2025 Jun 20;18(1):56. doi: 10.1186/s12284-025-00809-0. Rice (N Y). 2025. PMID: 40540085 Free PMC article. Review.
References
-
- Von Uexkull HR, Mutert E. 1995. Global extent, development and economic impact of acid soils. Plant Soil 171:1–15. doi:10.1007/BF00009558. - DOI
-
- Singh S, Tripathi DK, Singh S, Sharma S, Dubey NK, Chauhan DK, Vaculík M. 2017. Toxicity of aluminium on various levels of plant cells and organism: a review. Environ Exp Bot 137:177–193. doi:10.1016/j.envexpbot.2017.01.005. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

