The origin and evolution of a plant resistosome
- PMID: 35166827
- PMCID: PMC9048963
- DOI: 10.1093/plcell/koac053
The origin and evolution of a plant resistosome
Abstract
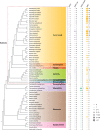
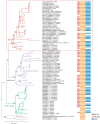
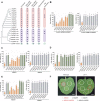
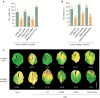
The nucleotide-binding, leucine-rich receptor (NLR) protein HOPZ-ACTIVATED RESISTANCE 1 (ZAR1), an immune receptor, interacts with HOPZ-ETI-DEFICIENT 1 (ZED1)-related kinases (ZRKs) and AVRPPHB SUSCEPTIBLE 1-like proteins to form a pentameric resistosome, triggering immune responses. Here, we show that ZAR1 emerged through gene duplication and that ZRKs were derived from the cell surface immune receptors wall-associated protein kinases (WAKs) through the loss of the extracellular domain before the split of eudicots and monocots during the Jurassic period. Many angiosperm ZAR1 orthologs, but not ZAR1 paralogs, are capable of oligomerization in the presence of AtZRKs and triggering cell death, suggesting that the functional ZAR1 resistosome might have originated during the early evolution of angiosperms. Surprisingly, inter-specific pairing of ZAR1 and AtZRKs sometimes results in the formation of a resistosome in the absence of pathogen stimulation, suggesting within-species compatibility between ZAR1 and ZRKs as a result of co-evolution. Numerous concerted losses of ZAR1 and ZRKs occurred in angiosperms, further supporting the ancient co-evolution between ZAR1 and ZRKs. Our findings provide insights into the origin of new plant immune surveillance networks.
© American Society of Plant Biologists 2022. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures




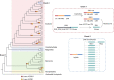
Comment in
-
Inventing the wheel: new insights into resistosome evolution.Plant Cell. 2022 Apr 26;34(5):1425-1426. doi: 10.1093/plcell/koac066. Plant Cell. 2022. PMID: 35201350 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

