Phosphorylation of MdERF17 by MdMPK4 promotes apple fruit peel degreening during light/dark transitions
- PMID: 35166845
- PMCID: PMC9048921
- DOI: 10.1093/plcell/koac049
Phosphorylation of MdERF17 by MdMPK4 promotes apple fruit peel degreening during light/dark transitions
Abstract
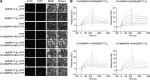
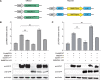
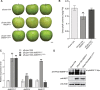
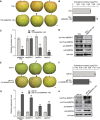
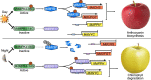
As apple fruits (Malus domestica) mature, they accumulate anthocyanins concomitantly with losing chlorophyll (Chl); however, the molecular pathways and events that coordinate Chl degradation and fruit coloration have not been elucidated. We showed previously that the transcription factor ETHYLENE RESPONSE FACTOR17 (MdERF17) modulates Chl degradation in apple fruit peels and that variation in the pattern of MdERF17 serine (Ser) residues is responsible for differences in its transcriptional regulatory activity. Here, we report that MdERF17 interacts with and is phosphorylated by MAP KINASE4 (MdMPK4-14G). Phosphorylation of MdERF17 at residue Thr67 by MdMPK4-14G is necessary for its transcriptional regulatory activity and its regulation of Chl degradation. We also show that MdERF17 mutants with different numbers of Ser repeat insertions exhibit altered phosphorylation profiles, with more repeats increasing its interaction with MdMPK4. MdMPK4-14G can be activated by exposure to darkness and is involved in the dark-induced degreening of fruit peels. We also demonstrate that greater phosphorylation of MdERF17 by MdMPK4-14G is responsible for the regulation of Chl degradation during light/dark transitions. Overall, our findings reveal the mechanism by which MdMPK4 controls fruit peel coloration.
© The Author(s) 2022. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Figures




Similar articles
-
Apple MPK4 mediates phosphorylation of MYB1 to enhance light-induced anthocyanin accumulation.Plant J. 2021 Jun;106(6):1728-1745. doi: 10.1111/tpj.15267. Epub 2021 May 7. Plant J. 2021. PMID: 33835607
-
Natural Variation Underlies Differences in ETHYLENE RESPONSE FACTOR17 Activity in Fruit Peel Degreening.Plant Physiol. 2018 Mar;176(3):2292-2304. doi: 10.1104/pp.17.01320. Epub 2018 Feb 5. Plant Physiol. 2018. PMID: 29431631 Free PMC article.
-
Low temperature delays degreening of apple fruit by inhibiting pheophorbide a oxygenase (PAO) pathway and chlorophyll oxidation during ripening.J Food Biochem. 2022 Aug;46(8):e14173. doi: 10.1111/jfbc.14173. Epub 2022 Apr 6. J Food Biochem. 2022. PMID: 35383957
-
Research progress of fruit color development in apple (Malus domestica Borkh.).Plant Physiol Biochem. 2021 May;162:267-279. doi: 10.1016/j.plaphy.2021.02.033. Epub 2021 Mar 4. Plant Physiol Biochem. 2021. PMID: 33711720 Review.
-
Regulatory Mechanisms of Anthocyanin Biosynthesis in Apple and Pear.Int J Mol Sci. 2021 Aug 6;22(16):8441. doi: 10.3390/ijms22168441. Int J Mol Sci. 2021. PMID: 34445149 Free PMC article. Review.
Cited by
-
Deciphering the regulatory network of the NAC transcription factor FvRIF, a key regulator of strawberry (Fragaria vesca) fruit ripening.Plant Cell. 2023 Oct 30;35(11):4020-4045. doi: 10.1093/plcell/koad210. Plant Cell. 2023. PMID: 37506031 Free PMC article.
-
Pan-genome analysis of 13 Malus accessions reveals structural and sequence variations associated with fruit traits.Nat Commun. 2023 Nov 15;14(1):7377. doi: 10.1038/s41467-023-43270-7. Nat Commun. 2023. PMID: 37968318 Free PMC article.
-
Spatial regulation of chlorophyll degradation in kiwifruit: AcNAC2-AcSGR1/2 cascades mediate rapid de-greening in the inner pericarp.Plant Biotechnol J. 2025 Jul;23(7):2554-2569. doi: 10.1111/pbi.70071. Epub 2025 Apr 4. Plant Biotechnol J. 2025. PMID: 40183233 Free PMC article.
-
Interaction of AcMADS68 with transcription factors regulates anthocyanin biosynthesis in red-fleshed kiwifruit.Hortic Res. 2022 Nov 15;10(2):uhac252. doi: 10.1093/hr/uhac252. eCollection 2023 Feb. Hortic Res. 2022. PMID: 36751270 Free PMC article.
-
PpERF17 alleviates peach fruit postharvest chilling injury under elevated CO2 by activating jasmonic acid and γ-aminobutyric acid biosynthesis.Hortic Res. 2025 Jan 15;12(4):uhaf014. doi: 10.1093/hr/uhaf014. eCollection 2025 Apr. Hortic Res. 2025. PMID: 40093381 Free PMC article.
References
-
- Allan AC, Hellens RP, Laing WA (2008) MYB transcription factors that colour our fruit. Trends Plant Sci 13: 99–102 - PubMed
-
- An X, Tian Y, Chen K, Liu X, Liu D, Xie X, Cheng C, Cong P, Hao Y (2015) MdMYB9 and MdMYB11 are involved in the regulation of the JA-induced biosynthesis of anthocyanin and proanthocyanidin in apples. Plant Cell Physiol 56: 650–662 - PubMed
-
- An X, Tian Y, Chen K, Wang X, Hao Y (2012) The apple WD40 protein MdTTG1 interacts with bHLH but not MYB proteins to regulate anthocyanin accumulation. J Plant Physiol 169: 710–717 - PubMed
-
- Andreasson E, Ellis B (2010) Convergence and specificity in the Arabidopsis MAPK nexus. Trends Plant Sci 15: 106–113 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

