Germinal centre-driven maturation of B cell response to mRNA vaccination
- PMID: 35168246
- PMCID: PMC9204750
- DOI: 10.1038/s41586-022-04527-1
Germinal centre-driven maturation of B cell response to mRNA vaccination
Abstract
Germinal centres (GC) are lymphoid structures in which B cells acquire affinity-enhancing somatic hypermutations (SHM), with surviving clones differentiating into memory B cells (MBCs) and long-lived bone marrow plasma cells1-5 (BMPCs). SARS-CoV-2 mRNA vaccination induces a persistent GC response that lasts for at least six months in humans6-8. The fate of responding GC B cells as well as the functional consequences of such persistence remain unknown. Here, we detected SARS-CoV-2 spike protein-specific MBCs in 42 individuals who had received two doses of the SARS-CoV-2 mRNA vaccine BNT162b2 six month earlier. Spike-specific IgG-secreting BMPCs were detected in 9 out of 11 participants. Using a combined approach of sequencing the B cell receptors of responding blood plasmablasts and MBCs, lymph node GC B cells and plasma cells and BMPCs from eight individuals and expression of the corresponding monoclonal antibodies, we tracked the evolution of 1,540 spike-specific B cell clones. On average, early blood spike-specific plasmablasts exhibited the lowest SHM frequencies. By contrast, SHM frequencies of spike-specific GC B cells increased by 3.5-fold within six months after vaccination. Spike-specific MBCs and BMPCs accumulated high levels of SHM, which corresponded with enhanced anti-spike antibody avidity in blood and enhanced affinity as well as neutralization capacity of BMPC-derived monoclonal antibodies. We report how the notable persistence of the GC reaction induced by SARS-CoV-2 mRNA vaccination in humans culminates in affinity-matured long-term antibody responses that potently neutralize the virus.
© 2022. The Author(s), under exclusive licence to Springer Nature Limited.
Figures


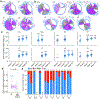





Update of
-
Germinal centre-driven maturation of B cell response to SARS-CoV-2 vaccination.bioRxiv [Preprint]. 2021 Nov 2:2021.10.31.466651. doi: 10.1101/2021.10.31.466651. bioRxiv. 2021. Update in: Nature. 2022 Apr;604(7904):141-145. doi: 10.1038/s41586-022-04527-1. PMID: 34751268 Free PMC article. Updated. Preprint.
Comment in
-
Severe acute respiratory syndrome coronavirus-2 vaccine-induced B cells aspire to long-lived connections: Tracking B-cell memory over time.Immunol Cell Biol. 2022 May;100(5):308-311. doi: 10.1111/imcb.12548. Epub 2022 Apr 16. Immunol Cell Biol. 2022. PMID: 35353930 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

