Lipid and Nucleocapsid N-Protein Accumulation in COVID-19 Patient Lung and Infected Cells
- PMID: 35171025
- PMCID: PMC8849100
- DOI: 10.1128/spectrum.01271-21
Lipid and Nucleocapsid N-Protein Accumulation in COVID-19 Patient Lung and Infected Cells
Abstract
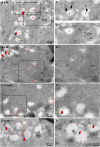
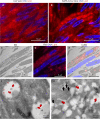
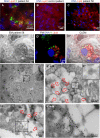
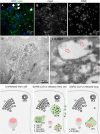
The pandemic of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused a global outbreak and prompted an enormous research effort. Still, the subcellular localization of the coronavirus in lungs of COVID-19 patients is not well understood. Here, the localization of the SARS-CoV-2 proteins is studied in postmortem lung material of COVID-19 patients and in SARS-CoV-2-infected Vero cells, processed identically. Correlative light and electron microscopy on semithick cryo-sections demonstrated induction of electron-lucent, lipid-filled compartments after SARS-CoV-2 infection in both lung and cell cultures. In lung tissue, the nonstructural protein 4 and the stable nucleocapsid N-protein were detected on these novel lipid-filled compartments. The induction of such lipid-filled compartments and the localization of the viral proteins in lung of patients with fatal COVID-19 may explain the extensive inflammatory response and provide a new hallmark for SARS-CoV-2 infection at the final, fatal stage of infection. IMPORTANCE Visualization of the subcellular localization of SARS-CoV-2 proteins in lung patient material of COVID-19 patients is important for the understanding of this new virus. We detected viral proteins in the context of the ultrastructure of infected cells and tissues and discovered that some viral proteins accumulate in novel, lipid-filled compartments. These structures are induced in Vero cells but, more importantly, also in lung of patients with COVID-19. We have characterized these lipid-filled compartments and determined that this is a novel, virus-induced structure. Immunogold labeling demonstrated that cellular markers, such as CD63 and lipid droplet marker PLIN-2, are absent. Colocalization of lipid-filled compartments with the stable N-protein and nonstructural protein 4 in lung of the last stages of COVID-19 indicates that these compartments play a key role in the devastating immune response that SARS-CoV-2 infections provoke.
Keywords: COVID-19; electron microscopy; lipids; pathology.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Chen J, Malone B, Llewellyn E, Grasso M, Shelton PMM, Olinares PDB, Maruthi K, Eng ET, Vatandaslar H, Chait BT, Kapoor TM, Darst SA, Campbell EA. 2020. Structural basis for helicase-polymerase coupling in the SARS-CoV-2 replication-transcription complex. Cell 182:1560–1573.e13. doi: 10.1016/j.cell.2020.07.033. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

