Comprehensive mapping of SARS-CoV-2 peptide epitopes for development of a highly sensitive serological test for total and neutralizing antibodies
- PMID: 35174857
- PMCID: PMC9005051
- DOI: 10.1093/protein/gzab033
Comprehensive mapping of SARS-CoV-2 peptide epitopes for development of a highly sensitive serological test for total and neutralizing antibodies
Abstract
Quantification of the anti-SARS-CoV-2 antibody response has proven to be a prominent diagnostic tool during the COVID-19 pandemic. Antibody measurements have aided in the determination of humoral protection following infection or vaccination and will likely be essential for predicting the prevalence of population level immunity over the next several years. Despite widespread use, current tests remain limited in part, because antibody capture is accomplished through the use of complete spike and nucleocapsid proteins that contain significant regions of overlap with common circulating coronaviruses. To address this limitation, a unique epitope display platform utilizing monovalent display and protease-driven capture of peptide epitopes was used to select high affinity peptides. A single round of selection using this strategy with COVID-19 positive patient plasma samples revealed surprising differences and specific patterns in the antigenicity of SARS-CoV-2 proteins, especially the spike protein. Putative epitopes were assayed for specificity with convalescent and control samples, and the individual binding kinetics of peptides were also determined. A subset of prioritized peptides was used to develop an antibody diagnostic assay that showed low cross reactivity while detecting 37% more positive antibody cases than a gold standard FDA EUA test. Finally, a subset of peptides were compared with serum neutralization activity to establish a 2 peptide assay that strongly correlates with neutralization. Together, these data demonstrate a novel phage display method that is capable of comprehensively and rapidly mapping patient viral antibody responses and selecting high affinity public epitopes for the diagnosis of humoral immunity.
Keywords: antibodies; epitope mapping; neutralizing; phage display; serological assay; serum ELISA.
© The Author(s) 2022. Published by Oxford University Press. All rights reserved. For Permissions, please e-mail: journals.permissions@oup.com.
Figures





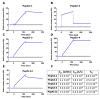
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

