The histone demethylase PHF8 regulates TGFβ signaling and promotes melanoma metastasis
- PMID: 35179962
- PMCID: PMC8856617
- DOI: 10.1126/sciadv.abi7127
The histone demethylase PHF8 regulates TGFβ signaling and promotes melanoma metastasis
Abstract
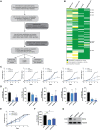
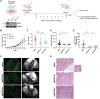
The contribution of epigenetic dysregulation to metastasis remains understudied. Through a meta-analysis of gene expression datasets followed by a mini-screen, we identified Plant Homeodomain Finger protein 8 (PHF8), a histone demethylase of the Jumonji C protein family, as a previously unidentified prometastatic gene in melanoma. Loss- and gain-of-function approaches demonstrate that PHF8 promotes cell invasion without affecting proliferation in vitro and increases dissemination but not subcutaneous tumor growth in vivo, thus supporting its specific contribution to the acquisition of metastatic potential. PHF8 requires its histone demethylase activity to enhance melanoma cell invasion. Transcriptomic and epigenomic analyses revealed that PHF8 orchestrates a molecular program that directly controls the TGFβ signaling pathway and, as a consequence, melanoma invasion and metastasis. Our findings bring a mechanistic understanding of epigenetic regulation of metastatic fitness in cancer, which may pave the way for improved therapeutic interventions.
Figures






References
-
- Chaffer C. L., Weinberg R. A., A perspective on cancer cell metastasis. Science 331, 1559–1564 (2011). - PubMed
-
- Bailey M. H., Tokheim C., Porta-Pardo E., Sengupta S., Bertrand D., Weerasinghe A., Colaprico A., Wendl M. C., Kim J., Reardon B., Ng P. K.-S., Jeong K. J., Cao S., Wang Z., Gao J., Gao Q., Wang F., Liu E. M., Mularoni L., Rubio-Perez C., Nagarajan N., Cortés-Ciriano I., Zhou D. C., Liang W.-W., Hess J. M., Yellapantula V. D., Tamborero D., Gonzalez-Perez A., Suphavilai C., Ko J. Y., Khurana E., Park P. J., Van Allen E. M., Liang H.; MC3 Working Group; Cancer Genome Atlas Research Network, Lawrence M. S., Godzik A., Lopez-Bigas N., Stuart J., Wheeler D., Getz G., Chen K., Lazar A. J., Mills G. B., Karchin R., Ding L., Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385.e18 (2018). - PMC - PubMed
-
- Hodis E., Watson I. R., Kryukov G. V., Arold S. T., Imielinski M., Theurillat J. P., Nickerson E., Auclair D., Li L., Place C., DiCara D., Ramos A. H., Lawrence M. S., Cibulskis K., Sivachenko A., Voet D., Saksena G., Stransky N., Onofrio R. C., Winckler W., Ardlie K., Wagle N., Wargo J., Chong K., Morton D. L., Stemke-Hale K., Chen G., Noble M., Meyerson M., Ladbury J. E., Davies M. A., Gershenwald J. E., Wagner S. N., Hoon D. S. B., Schadendorf D., Lander E. S., Gabriel S. B., Getz G., Garraway L. A., Chin L., A landscape of driver mutations in melanoma. Cell 150, 251–263 (2012). - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

