Virulence Biomarkers of Bursaphelenchus xylophilus: A Proteomic Approach
- PMID: 35211137
- PMCID: PMC8861294
- DOI: 10.3389/fpls.2021.822289
Virulence Biomarkers of Bursaphelenchus xylophilus: A Proteomic Approach
Abstract
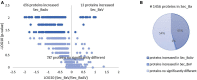
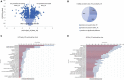
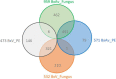
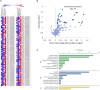
The pinewood nematode (PWN), Bursaphelenchus xylophilus, one of the most serious forest pests worldwide, is considered the causal agent of the pine wilt disease (PWD). The main host species belong to the genus Pinus, and a variation in the susceptibility of several pine species to PWN infection is well-known. It is also recognized that there is variation in the virulence among B. xylophilus isolates. In the present study, we applied a quantitative mass spectrometry-based proteomics approach to perform a deep characterization of proteomic changes across two B. xylophilus isolates with different virulence from different hosts and geographical origins. A total of 1,456 proteins were quantified and compared in the two isolates secretomes, and a total of 2,741 proteins were quantified and compared in the nematode proteomes in pine tree extract and fungus stimuli conditions. From the proteomic analyses, a group of proteins was selected and identified as potential virulence biomarkers and shed light on putative most pathogenic proteins of this plant-parasitic nematode. Proteomic data are available via ProteomeXchange with identifier PXD029377.
Keywords: pine trees; pine wilt disease; pinewood nematode; plant–nematode interactions; proteomics; secretome; virulence.
Copyright © 2022 Cardoso, Anjo, Manadas, Silva, Abrantes, Nakamura and Fonseca.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
-
- Aikawa T., Kikuchi T. (2007). Estimation of virulence of Bursaphelenchus xylophilus (Nematoda: Aphelenchoididae) based on its reproductive ability. Nematology 9 371–377.
-
- Aikawa T., Kikuchi T., Kosaka H. (2003a). Demonstration of interbreeding between virulent and avirulent populations of Bursaphelenchus xylophilus (Nematoda: Aphelenchoididae) by {PCR-RFLP} method. Appl. Entomol. Zool. 38 565–569. 10.1303/aez.2003.565 - DOI
-
- Aikawa T., Togashi K., Kosaka H. (2003b). Different developmental responses of virulent and avirulent isolates of the pinewood nematode, Bursaphelenchus xylophilus (Nematoda: Aphelenchoididae), to the insect vector, Monochamus alternatus (Coleoptera: Cerambycidae). Environ. Entomol. 32 96–102.
-
- Anjo S. I., Lourenço A. S., Melo M. N., Santa C., Manadas B. (2016). “Unraveling mesenchymal stem cells’ dynamic secretome through nontargeted proteomics profiling,” in Mesenchymal Stem Cells: Methods and Protocols, Vol. 1416 ed. Gnecchi M. (New York, NY: Human Press; ), 521–549. 10.1007/978-1-4939-3584-0_32 - DOI - PubMed
LinkOut - more resources
Full Text Sources

