Dermal αSMA+ myofibroblasts orchestrate skin wound repair via β1 integrin and independent of type I collagen production
- PMID: 35212000
- PMCID: PMC8982612
- DOI: 10.15252/embj.2021109470
Dermal αSMA+ myofibroblasts orchestrate skin wound repair via β1 integrin and independent of type I collagen production
Abstract
Skin wound repair is essential for organismal survival and failure of which leads to non-healing wounds, a leading health issue worldwide. However, mechanistic understanding of chronic wounds remains a major challenge due to lack of appropriate genetic mouse models. αSMA+ myofibroblasts, a unique class of dermal fibroblasts, are associated with cutaneous wound healing but their precise function remains unknown. We demonstrate that genetic depletion of αSMA+ myofibroblasts leads to pleiotropic wound healing defects, including lack of reepithelialization and granulation, dampened angiogenesis, and heightened hypoxia, hallmarks of chronic non-healing wounds. Other wound-associated FAP+ and FSP1+ fibroblasts do not exhibit such dominant functions. While type I collagen (COL1) expressing cells play a role in the repair process, COL1 produced by αSMA+ myofibroblasts is surprisingly dispensable for wound repair. In contrast, we show that β1 integrin from αSMA+ myofibroblasts, but not TGFβRII, is essential for wound healing, facilitating contractility, reepithelization, and vascularization. Collectively, our study provides evidence for the functions of myofibroblasts in β1 integrin-mediated wound repair with potential implications for treating chronic non-healing wounds.
Keywords: extracellular matrix; myofibroblasts; wound healing.
© 2022 The Authors.
Figures
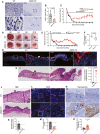
- A–E
Impaired wound closure in αSMA‐TK mice. (A) IHC for αSMA+ cells in wound tissue sections at day 6 post‐wounding. Representative images are shown. Scale bar, 50 µm. Black boxes denote zoomed regions displayed in bottom panels. (B) Quantification of αSMA+ cells at day 6 post‐wounding. WT, N = 6; TK, N = 5 biological replicates. Two‐tailed non‐parametric Mann–Whitney test performed. (C) WT and αSMA‐TK mice were treated with GCV for 32 days. Wounds were measured at indicated timepoints and wound area is presented as percentage of day 0 value. WT, N = 5; TK, N = 4 biological replicates per group, data are from one experiment. Two‐way ANOVA performed comparing WT to TK mice. (D) Representative gross images of wounds. Scale bar, 5 mm. (E) αSMA‐TK mice and WT littermates were placed on GCV regimen 3 days prior to wounding (day −3). On day 21, GCV treatment was stopped and the mice monitored until day 39. WT, N = 7; TK, N = 9 biological replicates. Data are from two independent experiments. Two‐way ANOVA and Sidak’s multiple comparison test were performed comparing WT to TK mice.
- F
Expression levels of ACTA2 in normal and diabetic human wound tissue. RNA‐seq reads are derived from Davis et al (2020c), Data ref: Davis et al (2020a). Normal, N = 3; diabetic, N = 4 biological replicates. Data are presented as a box and whisker plot of min to max values. Unpaired t‐test performed.
- G–I
Impaired re‐epithelialization of the wounds in αSMA‐TK mice at day 17 post‐wounding. (G) Representative images of keratin 5 (Krt5, red) and Hoechst (blue) stained wound sections at day 17 post‐wounding. White boxes denote zoomed regions displayed in right panels, triangles (open: incomplete closure, filled: closed wounds) denote the epithelial tongues. Scale bar, 1,000 µm. (H) Representative H&E images of wound sections. Wound bed is marked with black lines, triangles (open: incomplete closure, filled: closed wounds) denote the epithelial tongues. Epi, epidermis; derm, dermis; subq, subcutaneous regions denoted. Scale bar, 200 µm. (I) Measurements of the distances between epithelial tips in the wounded skins of WT and αSMA‐TK mice at day 17 post‐wounding. WT, N = 7; TK, N = 8 biological replicates. Black line denotes median value. Mann–Whitney test performed.
- J–O
Impaired granulation tissue formation and angiogenesis in αSMA‐TK mice. (J) Representative H&E images of wound tissue sections at day 17 post‐wounding. Granulation areas are marked by black lines, triangles (open: incomplete closure, filled: closed wounds) denote the epithelial tongues. Epi, epidermis; derm, dermis regions denoted. Scale bar: 200 μm. (K) Granulation tissue thickness measured on digital images and normalized to the average values in the WT control (set to 100%). NA (not assessed) reflects the lack of granulation in αSMA‐TK mice. WT, N = 5; TK, N = 4 biological replicates. (L) Immunofluorescent staining for CD31 (red) and nuclei (blue) on wound tissue sections at day 17 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar, 100 μm. (M) Quantification of CD31+ cells per visual field normalized to WT mice. WT, N = 3; TK, N = 5 biological replicates. (N) IHC for hypoxyprobe (brown) on wound tissue sections at day 17 post‐wounding. Black boxes denote zoomed regions depicted in right panels. Scale bar, 100 μm. (O) Quantification of hypoxyprobe area per visual field normalized to WT mice. WT, N = 5; TK, N = 6 biological replicates.

Representative images of wound tissue at day 17 post‐wounding stained for αSMA (green), CD31 (red), and Hoechst (blue). Scale bar, 20 μm. White boxes denote zoomed regions displayed in bottom panels.
Quantification of CD31+αSMA+ cells in wound tissue at day 17 post‐wounding. WT, N = 5; TK, N = 5 biological replicates.
Representative images of hair follicles in normal skin surrounding wound tissue at day 17 post‐wounding stained for αSMA (green) and Hoechst (blue). Scale bar, 20 μm.
Quantification of αSMA+ cells in normal skin surrounding wound tissue at day 17 post‐wounding. WT, N = 5; TK, N = 6 biological replicates.
Representative images of liver, kidney, and lung stained for αSMA (green), CD31 (red), and Hoechst (blue). Scale bar, 20 μm.
Quantification of αSMA+ cells in liver, kidney, and lung at day 17 post‐wounding. Liver: WT, N = 6; TK, N = 6 biological replicates. Kidney: WT, N = 7; TK, N = 7 biological replicates. Lung: WT, N = 7; TK, N = 6 biological replicates.
- A
Representative images of wound tissue at day 41 post‐wounding stained for αSMA (red) and Hoechst (blue) from mice treated with GCV initiated at 3 days prior to wounding and continued until 21 days post‐wounding followed by withdrawal of GCV treatment (related to Fig 1E). White boxes denote zoomed regions displayed in right panels. Scale bar, 100 µm.
- B, C
Impaired re‐epithelialization of the wounds in αSMA‐TK mice at day 6 post‐wounding. (B) Representative images of keratin 5 (Krt5, red) and Hoechst (blue) stained wound sections. White boxes denote zoomed regions displayed in right panels. Scale bar, 1,000 µm. (C) Representative H&E images of wound sections. Wound area is marked with black dashed lines, open triangles denote the epithelial tongues. Epi, epidermis; derm, dermis; subq, subcutaneous regions denoted. Scale bar, 200 µm.
- D
Measurements of the distances between epithelial tips in the wounded skins of WT and αSMA‐TK mice at day 6 post‐wounding. WT, N = 6; TK, N = 7 biological replicates. Black lines denote median values. Unpaired Student’s t‐test performed.
- E, F
Impaired granulation tissue formation in αSMA‐TK mice at day 6 post‐wounding. (E) Representative H&E images of wound tissue sections at day 6 post‐wounding. Granulation areas are marked by black lines. Derm, dermis and subq, subcutaneous regions denoted. Scale bar, 500 μm. (F) Granulation tissue thickness measured on digital images and normalized to the average values in the WT control (set to 100%). NA (not assessed) reflects the lack of granulation in αSMA‐TK mice. WT, N = 5; TK, N = 7 biological replicates.
- G
Representative IHC images of day 17 wound tissue stained with Gr1 antibody. Scale bar, 50 μm.
- H
Quantitative analysis of Gr1+ area in day 17 wound tissue. WT, N = 6; TK, N = 7 biological replicates. Two‐tailed Mann–Whitney test performed comparing WT to TK.
- I
Unsupervised clustering of gene expression of day 6 wounds, when the highest numbers of αSMA+ fibroblasts are observed, and day 17, when the wound closure is achieved in WT control mice. The main gene categories altered in αSMA‐TK mice are marked on the right. For day 6 wounds, N = 3 biological replicates per group were included, and each sample was analyzed in technical duplicates. Day 17 analysis was performed with N = 2 biological replicates per group with each sample analyzed in technical triplicates.

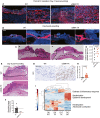
- A
Analysis of multiplex immunostaining of full‐thickness skin wounds performed at indicated time points. Data are presented as the fractional area of each image positive for αSMA and negative for CD31. Day 0, N = 2; Day 6, N = 4; Day 9, N = 3; Day 14, N = 3; Day 21, N = 3; Day 32, N = 4 biological replicates. One‐way ANOVA with Dunnett’s multiple comparison test performed.
- B
FACS quantification of αSMA‐RFP+ cells in wounds. Day 0, N = 2; day 3, N = 2; day 6, N = 6; day 8, N = 3; day 14, N = 3 biological replicates. One‐way ANOVA with Dunnett’s multiple comparison test performed comparing each timepoint to day 0.
- C
Schematic of bone marrow transplants between WT and αSMA‐RFP mice.
- D
Representative FACS plots of digested skin from bone marrow (BM) transplanted αSMA‐RFP mice 17 days post‐wounding.
- E
FACS quantification of αSMA‐RFP+ cells in wounds of BM transplanted mice. αSMA‐RFP with αSMA‐RFP BM, N = 2; WT with αSMA‐RFP BM, N = 3; αSMA‐RFP with WT BM, N = 4 biological replicates. One‐way ANOVA with Tukey’s multiple comparison test performed comparing WT with αSMA‐RFP BM and αSMA‐RFP with WT BM to αSMA‐RFP with αSMA‐RFP BM.
- F
Schematic of bone marrow transplants between WT and αSMA‐TK mice.
- G
Wound area measurements in WT and ⍺SMA‐TK mice transplanted with WT or αSMA‐TK BM. WT with WT BM and WT with αSMA‐TK BM, N = 5; αSMA‐TK with WT BM and αSMA‐TK with αSMA‐TK BM, N = 4 biological replicates. Two‐way ANOVA performed with the indicated comparisons.
- H–J
Multispectral imaging of changes in fibroblast subsets during wound repair. (H) Representative images (20× magnification) of orthogonal wound tissue sections at day 6 post‐wounding displaying αSMA, FAP, and FSP1 overlaid with DAPI in the wound bed. Subcutaneous (subq), dermis (derm), epidermis (epi), and scar regions are denoted. Individual αSMA, FAP, and FSP1 channel images overlaid with DAPI. Scale bar, 50 µm. (I) Analysis of multiplex immunostaining of full‐thickness skin wounds performed at indicated time points. Data are presented as the fractional area of each image positive for the indicated markers and negative for CD31. Day 0, N = 2; Day 6, N = 4; Day 9, N = 3; Day 14, N = 3; Day 21, N = 3; Day 32, N = 4 biological replicates. One‐way ANOVA with Dunnett’s multiple comparison test performed. (J) Percent overlap of αSMA with FAP and FSP1 throughout wound healing. αSMA+ denotes cells positive for αSMA but not FAP or FSP1. Day 0, N = 2; Day 6, N = 4; Day 9, N = 3; Day 14, N = 3; Day 21, N = 3; Day 32, N = 4 biological replicates.
- K
Wound closure in FAP‐TK and wild‐type littermate mice. N = 4 biological replicates per group, data are from one experiment. Two‐way ANOVA performed comparing WT to TK mice.
- L
Representative H&E images of wound tissue sections at day 17 post‐wounding. Granulation areas are marked by black lines. Scale bar, 200 μm.
- M
Granulation tissue thickness measured on digital images and normalized to the average values in the WT control. WT, N = 3; FAP‐TK, N = 3; WT, N = 17, FSP1‐TK, N = 9 biological replicates.
- N
Immunofluorescent staining for CD31 (red) and nuclei on wound tissue sections at day 17 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar, 100 μm.
- O
Quantification of CD31+ cells per visual field normalized to WT mice. WT, N = 4; FAP‐TK, N = 4; WT, N = 3; FSP1‐TK, N = 3 biological replicates.
- P
IHC for hypoxyprobe (brown) on wound tissue sections at day 17 post‐wounding. Black boxes denote zoomed regions depicted in right panels. Scale bar, 100 μm.
- Q
Quantification of hypoxyprobe area per visual field normalized to WT mice. WT, N = 3; FAP‐TK, N = 3; WT, N = 3; FSP1‐TK, N = 4 biological replicates.
- R
Wound closure in FSP1‐TK and WT littermate mice. N = 10 biological replicates per group, data are from two independent experiments. Two‐way ANOVA performed comparing WT to TK mice.

FACS gating strategy for Fig 2B. αSMA‐RFP‐ control: skin tissue from an αSMA‐RFP‐ (WT) mouse.
Representative Y‐chromosome (red) fluorescent in situ hybridization (FISH) images of bone marrow cells after bone marrow transplantation. White boxes denote zoomed regions depicted in bottom panels. Negative control: secondary antibody alone stained tissue. Scale bar, 50 μm.
FACS gating strategy for Fig 2D and E. αSMA‐RFP‐ control: skin tissue from an αSMA‐RFP‐ (WT) mouse.
Expression level of putative fibroblast marker genes in normal and diabetic human wound tissue. RNA‐seq reads are derived from Davis et al (2020c), Data ref: Davis et al (2020a). Normal, N = 3; diabetic, N = 4 biological replicates. Data are presented as a box and whisker plot of min to max values. PDGFRA, TGFBR2, DPP4, VIM: Mann–Whitney test performed; PRRX1, THY1, FAP: Welch’s t‐test performed; EN1, S100A4, DDR2: unpaired t‐test performed.
Construct for newly generated FAP‐TK transgenic mice used in the study (left panel). The transgenic cassette contains the promoter and exon 1 regions (extended promoter) of FAP followed by HSV thymidine kinase and SV40 polyadenylation site. Dose‐dependent ablation of cultured fibroblasts from the ears of FAP‐TK mice and their wild‐type littermates (right panel). Data are normalized to the viability at 0 μM. WT, N = 2; TK, N = 5 biological replicates. Two‐way ANOVA with Sidak’s multiple comparison test performed comparing WT to TK.
Representative IHC images of FAP in day 17 wound tissue. Black boxes denote zoomed regions depicted in bottom panels. Scale bar, 50 μm.
Quantification of FAP+ cells in day 17 wound tissue. N = 5 biological replicates per group. Two‐tailed non‐parametric Mann–Whitney test performed comparing WT to TK mice.
Representative IHC images of FSP1 in day 17 wound tissue. Black boxes denote zoomed regions depicted in bottom panels. Scale bar, 50 µm.
Quantification of FSP1+ cells in day 17 wound tissue. WT, N = 8; TK+, N = 6 biological replicates. Two‐tailed non‐parametric Mann–Whitney test performed comparing WT to TK mice.
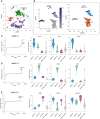
UMAP plot defining cell clusters. Expression of defining markers for each cluster is presented in Appendix Fig S1A.
UMAP plot of Acta2 (encoding αSMA) expression in the fibroblast cluster (left panel). UMAP plot of fibroblast subclusters (right panel).
Enrichment plot of genes in αSMA C1 (left panel). Violin plots for genes enriched in αSMA C1 (right panels).
Enrichment plot of genes in αSMA C2 (left panel). Violin plots for genes enriched in αSMA C2 (right panels).
Enrichment plot of genes in αSMA C3 (left panel). Violin plots of enriched genes in the αSMA C3 cluster (right panels). Raw scRNA‐seq data are derived from Haensel et al (2020b), Data ref: Haensel et al (2020a).

- A
IHC for collagen I on wound tissue sections at day 17 post‐wounding (brown stain). Black boxes denote zoomed regions depicted in bottom panels. Scale bar: 50 μm.
- B
Quantification of collagen I‐positive area normalized to the average area in WT controls. WT, N = 5; TK, N = 5 biological replicates.
- C
Polarized light images of picrosirius red stained wound tissues at day 17 post‐wounding. Red: collagen fibers. White boxes denote zoomed regions depicted in bottom panels. Scale bar, 50 µm.
- D
Quantification of collagen fiber (picrosirius red) percent area normalized to the average area in WT controls. WT, N = 3; TK, N = 3 biological replicates.
- E–H
Reduced collagen deposition in Col1α1cKO wounds. (E) IHC for collagen I on wound tissue sections at day 14 post‐wounding (brown stain). Black boxes denote zoomed regions depicted in bottom panels. Scale bar: 25 μm. (F) Quantification of collagen I‐positive area normalized to the average area in WT controls. WT, N = 7; Col1α1cKO, N = 4 biological replicates. (G) Polarized light images of picrosirius red stained wound tissues at day 14 post‐wounding. Red: collagen fibers. White boxes denote zoomed regions depicted in bottom panels. Scale bar, 50 µm. (H) Quantification of collagen fiber percent area and fiber width normalized to the average area and width, respectively, in WT controls. WT, N = 4; Col1α1cKO, N = 5 biological replicates.
- I
Wound area measurements in WT and Col1α1cKO mice. WT, N = 7; Col1α1cKO, N = 5 biological replicates. Data are from two independent experiments. Two‐way ANOVA performed comparing WT and Col1α1cKO mice.
- J–M
Reepithelization and vascularization in WT and Col1α1cKO mice. (J) Representative images of keratin 5 (Krt5, red) stained wounds at day 14 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar, 500 µm. (K) Quantification of epidermal thickness. WT, N = 6; Col1α1cKO, N = 5 biological replicates. (L) Immunofluorescent staining for CD31 (red) and nuclei (blue) on wound tissue sections at day 14 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar, 20 μm. (M) Quantification of CD31+ area per visual field normalized to WT mice. WT, N = 6; Col1α1cKO, N = 5 biological replicates.
- N
UMAP (left panel) and violin plots (right panel) of Col1a1 expression in murine wound fibroblasts. Raw scRNA‐seq data are derived from Haensel et al (2020b), Data ref: Haensel et al (2020a).
- O
Wound area measurements in Col1α1‐TK and WT littermate mice. WT, N = 13; Col1α1‐TK, N = 12 biological replicates, data are from two independent experiments. Two‐way ANOVA performed comparing WT to TK mice.
- P
Granulation tissue thickness measured on digital images and normalized to the average values in the WT control. WT, N = 4; TK, N = 7 biological replicates.

- A
Fold change in mRNA expression of Col1a1 in wound tissue. Expression is normalized to day 0 (D0, unwounded skin) values. Day 0 WT, Day 3 WT, Day 3 αSMA‐TK, and Day 6 αSMA‐TK, N = 6 mice; Day 6 WT, Day 17 WT, and Day 17 αSMA‐TK, N = 5 biological replicates. Unpaired t‐test comparing WT to αSMA‐TK for each time point performed based on ΔCT values.
- B, C
Assessment of the mechanical properties of wound sections from WT and αSMA‐TK mice at day 6 post‐wounding. N = 3 biological replicates per group, four positions per wound, 257 measurements per position. (B) Representative elasticity maps. The color map indicates the modulus of elasticity in kPa evaluated using force curves generated for each point in the field. This measurement provides AFM heights relative to the lowest contact point in the field. (C) Distribution of the Young’s modulus throughout the samples and the median values indicated on each histogram.
- D
Representative image (20× magnification) at day 6 post‐wounding displaying αSMA (red), FAP (orange), FSP1 (teal), and pro‐collagen I (green) overlaid with DAPI (blue, left panel). Individual αSMA, FAP, FSP1, and pro‐collagen I channel images overlaid with DAPI (right panel). Scale bar: 50 µm.
- E
Analysis of multiplex immunostaining of full‐thickness skin wounds performed at indicated time points. Data are presented as the fractional area of each image positive for a given marker and negative for CD31. Day 0, N = 2; Day 6, N = 4; Day 9, N = 3; Day 14, N = 3; Day 21, N = 3; Day 32, N = 4 biological replicates.
- F
Percent overlap of pro‐collagen I with other markers throughout wound healing. Pro‐collagen I+ denotes cells positive for pro‐collagen I but not αSMA, FAP, or FSP1.
- G
Construct for newly generated Col1α1‐TK transgenic mice used in the study. The transgenic cassette contains the promoter and exon 1 regions (extended promoter) of Col1a1 followed by HSV thymidine kinase and SV40 polyadenylation site.
- H
Dose‐dependent ablation of cultured fibroblasts from the ears of Col1α1‐TK mice and their wild‐type littermates. WT, N = 3; Col1α1‐TK+, N = 3 biological replicates. Two‐way ANOVA with Sidak’s multiple comparison test performed comparing WT to TK.
- I
Immunofluorescent images of ear fibroblasts from Col1α1‐TK mice stained for collagen I (red, top panels)) and αSMA (red, bottom panels) after GCV or vehicle (PBS) treatment for 5 days. Scale bar, 100 μm.
- J
Representative images of αSMA staining (red) and Col1a1 ISH (black) in day 17 WT wound tissue. Scale bar, 100 μm.
- K
Representative IHC images of collagen I in day 17 wound tissue. Black boxes denote zoomed regions depicted in bottom panels. Scale bar, 200 μm.
- L
Quantification of collagen I intensity in day 17 wound tissue. Data are normalized to collagen I intensity of unwounded skin. WT, N = 3; Col1α1‐TK+, N = 6 biological replicates. Two‐tailed non‐parametric Mann–Whitney test performed comparing WT to TK.

- A
Violin plot of Itgb1 expression levels in fibroblasts from murine wounds. Raw scRNA‐seq data are derived from Haensel et al (2020b), Data ref: Haensel et al (2020a).
- B
UMAP plots of ACTA2 and ITGB1 expression of fibroblasts in human wounds. Raw scRNA‐data are derived from Davis et al (2020c), Data ref: Davis et al (2020b).
- C
IF for β1 integrin (red) and αSMA (green) on wound tissue sections at day 17 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar: 20 μm.
- D
Quantification of the number of αSMA+ β1 integrin+ cells. WT, N = 3; TK, N = 3 biological replicates. Unpaired t‐test performed comparing WT to TK.
- E
Wound area measurements in WT and Itgb1cKO mice. WT, N = 5; Itgb1cKO, N = 6 biological replicates. Two‐way ANOVA performed comparing WT and Itgb1cKO mice.
- F–K
Reepithelization, granulation tissue formation, and vascularization in WT and Itgb1cKO mice. (F) Representative images of keratin 5 (Krt5, red) stained wounds at day 14 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar, 500 µm. (G) Quantification of epidermal thickness. WT, N = 4; Itgb1cKO, N = 6 biological replicates. (H) Immunofluorescent staining for CD31 (red) and nuclei (blue) on wound tissue sections at day 14 post‐wounding. White boxes denote zoomed regions depicted in right panels. Scale bar, 20 μm. (I) Quantification of CD31+ area per visual field normalized to WT mice. WT, N = 6; Itgb1cKO, N = 6 biological replicates. (J) Polarized light images of picrosirius red stained wound tissues at day 14 post‐wounding. Red: collagen fibers. White boxes denote zoomed regions depicted in right panels. Scale bar, 50 µm. (K) Quantification of collagen fiber percent area normalized to the average area in WT controls. WT, N = 6; Itgb1cKO, N = 6 biological replicates.
- L
Quantification of collagen fiber width normalized to the average width in WT controls. WT, N = 6; Itgb1cKO, N = 6 biological replicates.

- A
IHC for αSMA in day 17 wound tissue. Representative images are shown. Scale bar, 25 µm.
- B
Quantification of αSMA+ area in day 17 wound tissue per visual field normalized to WT mice. WT, N = 5; Itgb1cKO, N = 4 biological replicates. One sample t‐test performed comparing values to WT average.
- C
UMAP plot of Tgfbr2 in fibroblasts in murine wounds (left panel). Violin plot of Tgfbr2 levels in each fibroblast cluster (right panel). Raw scRNA‐seq data are derived from Haensel et al (2020b), Data ref: Haensel et al (2020a).
- D, E
Representative H&E (D) and Masson’s trichrome (E) images of wound sections of WT and Tgfbr2cKO mice at day 17 post‐wounding. Scale bar, 100 μm.
- F
Wound closure in WT and Tgfbr2cKO mice. N = 9 biological replicates per group, data are from two independent experiments. Two‐way ANOVA performed comparing WT to Tgfbr2cKO mice.
References
-
- Becht E, McInnes L, Healy J, Dutertre CA, Kwok IWH, Ng LG, Ginhoux F, Newell EW (2018) Dimensionality reduction for visualizing single‐cell data using UMAP. Nat Biotechnol 37: 38–44. - PubMed
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

