Advances in the Molecular Cytogenetics of Bananas, Family Musaceae
- PMID: 35214815
- PMCID: PMC8879896
- DOI: 10.3390/plants11040482
Advances in the Molecular Cytogenetics of Bananas, Family Musaceae
Abstract
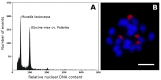
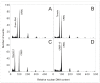
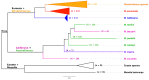
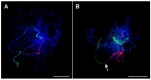
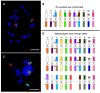
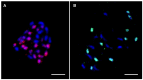
The banana is a staple food crop and represents an important trade commodity for millions of people living in tropical and subtropical countries. The most important edible banana clones originated from natural crosses between diploid Musa balbisiana and various subspecies of M. acuminata. It is worth mentioning that evolution and speciation in the Musaceae family were accompanied by large-scale chromosome structural changes, indicating possible reasons for lower fertility or complete sterility of these vegetatively propagated clones. Chromosomal changes, often accompanied by changes in genome size, are one of the driving forces underlying speciation in plants. They can clarify the genomic constitution of edible bananas and shed light on their origin and on diversification processes in members of the Musaceae family. This article reviews the development of molecular cytogenetic approaches, ranging from classical fluorescence in situ hybridization (FISH) using common cytogenetic markers to oligo painting FISH. We discuss differences in genome size and chromosome number across the Musaceae family in addition to the development of new chromosome-specific cytogenetic probes and their use in genome structure and comparative karyotype analysis. The impact of these methodological advances on our knowledge of Musa genome evolution at the chromosomal level is demonstrated. In addition to citing published results, we include our own new unpublished results and outline future applications of molecular cytogenetics in banana research.
Keywords: BAC clones; DNA repeats; chromosomes; flow cytometry; fluorescence in situ hybridization; karyotyping; oligo painting; rRNA genes.
Conflict of interest statement
All authors declare no conflict of interest.
Figures


Similar articles
-
Chromosome Painting Facilitates Anchoring Reference Genome Sequence to Chromosomes In Situ and Integrated Karyotyping in Banana (Musa Spp.).Front Plant Sci. 2019 Nov 20;10:1503. doi: 10.3389/fpls.2019.01503. eCollection 2019. Front Plant Sci. 2019. PMID: 31824534 Free PMC article.
-
Chromosome Painting in Cultivated Bananas and Their Wild Relatives (Musa spp.) Reveals Differences in Chromosome Structure.Int J Mol Sci. 2020 Oct 24;21(21):7915. doi: 10.3390/ijms21217915. Int J Mol Sci. 2020. PMID: 33114462 Free PMC article.
-
Striking variation in chromosome structure within Musa acuminata subspecies, diploid cultivars, and F1 diploid hybrids.Front Plant Sci. 2024 Jul 4;15:1387055. doi: 10.3389/fpls.2024.1387055. eCollection 2024. Front Plant Sci. 2024. PMID: 39027673 Free PMC article.
-
Oligonucleotide Fluorescence In Situ Hybridization: An Efficient Chromosome Painting Method in Plants.Plants (Basel). 2023 Jul 29;12(15):2816. doi: 10.3390/plants12152816. Plants (Basel). 2023. PMID: 37570972 Free PMC article. Review.
-
Fluorescence in situ hybridization in plants: recent developments and future applications.Chromosome Res. 2019 Sep;27(3):153-165. doi: 10.1007/s10577-019-09607-z. Epub 2019 Mar 9. Chromosome Res. 2019. PMID: 30852707 Review.
Cited by
-
A Flow Cytometry-Based Assessment of the Genomic Size and Ploidy Level of Wild Musa Species in India.Plants (Basel). 2023 Oct 18;12(20):3605. doi: 10.3390/plants12203605. Plants (Basel). 2023. PMID: 37896068 Free PMC article.
-
Characterizing subgenome recombination and chromosomal imbalances in banana varietal lineages.Ann Bot. 2024 Apr 10;133(2):349-364. doi: 10.1093/aob/mcad192. Ann Bot. 2024. PMID: 38097270 Free PMC article.
-
Unravelling genomic drivers of speciation in Musa through genome assemblies of wild banana ancestors.Nat Commun. 2025 Jan 23;16(1):961. doi: 10.1038/s41467-025-56329-4. Nat Commun. 2025. PMID: 39843949 Free PMC article.
References
-
- Häkkinen M. Reappraisal of sectional taxonomy in Musa (Musaceae) Taxon. 2013;62:809–813. doi: 10.12705/624.3. - DOI
-
- Janssens S.B., Vandelook F., de Langhe E., Verstraete B., Smets E., van den Houwe I., Swennen R. Evolutionary dynamics and biogeography of Musaceae reveal a correlation between the diversification of the banana family and the geological and climatic history of Southeast Asia. New Phytol. 2016;210:1453–1465. doi: 10.1111/nph.13856. - DOI - PMC - PubMed
-
- FAOSTAT Agriculture Organization of the United Nations, FAO. 2019. [(accessed on 5 June 2021)]. Available online: http://www.fao.org/home/en/
-
- Cheesman E.E. Classification of the bananas. The genus Ensete Horan. Kew Bull. 1947;2:97–117. doi: 10.2307/4109206. - DOI
-
- International Plant Genetic Resources Institute-International Network for the Improvement of Banana and Plantain/Centre de Coopération internationale en recherche agronomique pour le développement [IPGRI-INIBAP/CIRAD] Description for Banana (Musa spp.) Int. Network for the Improvement of Banana and Plantain; Montpellier, France: Centre de coopération int. en recherche agronomique pour le développement; Montpellier, France: International Plant Genetic Resources Institute Press; Rome, Italy: 1996.
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

