Features of Chromosome Introgression from Gossypium barbadense L. into G. hirsutum L. during the Development of Alien Substitution Lines
- PMID: 35214875
- PMCID: PMC8877206
- DOI: 10.3390/plants11040542
Features of Chromosome Introgression from Gossypium barbadense L. into G. hirsutum L. during the Development of Alien Substitution Lines
Abstract
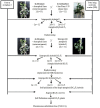
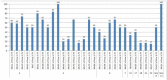
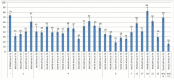
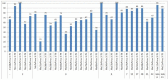
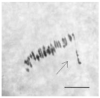
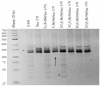
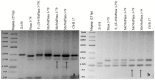
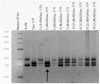
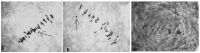
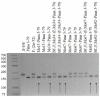
The creation of G. barbadense L./G. hirsutum L. chromosome-substitution lines is an important method to transfer agronomically valuable traits from G. barbadense into G. hirsutum. In this study, 30 monosomic lines of G. hirsutum from the Cytogenetic Collection of Uzbekistan, created in the genotypic background of line L-458, were used in crosses with the G. barbadense line Pima 3-79 to create substitution lines. In the course of this work, new monosomic lines were identified for chromosome 12 and monotelodisome 6 of the Atsubgenome and for chromosomes 17, 21, and 22 of the Dtsubgenome using chromosome-specific SSR markers and a well-defined tester set of cotton translocation lines (USA). Compared to those in the F1 hybrids, a strong decrease in the crossing and setting rates was found in the BC1F1 backcross lines, with the substitution of chromosomes 2, 4, 6, 7, and 12 of the Atsubgenome and 17, 18, 21, and 22 of the Dtsubgenome. The F1 and BC1F1 offspring from interspecific crosses differed in their transmission of univalents. Despite the regular pairing of chromosomes and the high meiotic index, interspecific aneuploid hybrids were characterized by a decrease in pollen fertility, which may indicate hidden structural variability in these genomes that did not affect meiotic division. The identification of chromosomes using chromosome-specific SSR markers in the early stages of plant development has greatly accelerated the detection of monosomic plants. The analysis of morphobiological traits revealed that monosomic F1 hybrids were more similar to the donor line, while BC1F1 hybrids were more similar to the recurrent parent but also showed previously undetected traits.
Keywords: G. barbadense; G. hirsutum; chromosome substitution lines; cotton; molecular markers; monosomic lines; translocation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

























Similar articles
-
Molecular-genetic and cytogenetic analyses of cotton chromosome introgression from Gossypium barbadense L. into the genome of G. hirsutum L. in BC2F1 hybrids.Vavilovskii Zhurnal Genet Selektsii. 2023 Dec;27(8):958-970. doi: 10.18699/VJGB-23-110. Vavilovskii Zhurnal Genet Selektsii. 2023. PMID: 38239965 Free PMC article.
-
Delineation of interspecific epistasis on fiber quality traits in Gossypium hirsutum by ADAA analysis of intermated G. barbadense chromosome substitution lines.Theor Appl Genet. 2011 May;122(7):1351-61. doi: 10.1007/s00122-011-1536-5. Epub 2011 Feb 8. Theor Appl Genet. 2011. PMID: 21301803
-
Construction of a complete set of alien chromosome addition lines from Gossypium australe in Gossypium hirsutum: morphological, cytological, and genotypic characterization.Theor Appl Genet. 2014 May;127(5):1105-21. doi: 10.1007/s00122-014-2283-1. Epub 2014 Feb 20. Theor Appl Genet. 2014. PMID: 24553965 Free PMC article.
-
Interspecific chromosomal effects on agronomic traits in Gossypium hirsutum by AD analysis using intermated G. barbadense chromosome substitution lines.Theor Appl Genet. 2013 Jan;126(1):109-17. doi: 10.1007/s00122-012-1965-9. Epub 2012 Sep 4. Theor Appl Genet. 2013. PMID: 22945267
-
Interspecific and intergeneric hybridization and chromosomal engineering of Brassicaceae crops.Breed Sci. 2014 May;64(1):14-22. doi: 10.1270/jsbbs.64.14. Breed Sci. 2014. PMID: 24987287 Free PMC article. Review.
Cited by
-
Genetic Analysis of Mutagenesis That Induces the Photoperiod Insensitivity of Wild Cotton Gossypium hirsutum Subsp. purpurascens.Plants (Basel). 2022 Nov 8;11(22):3012. doi: 10.3390/plants11223012. Plants (Basel). 2022. PMID: 36432741 Free PMC article.
-
Exploring agro-morphological and fiber traits diversity in cotton (G. barbadense L.).BMC Plant Biol. 2024 May 15;24(1):403. doi: 10.1186/s12870-024-04912-0. BMC Plant Biol. 2024. PMID: 38750434 Free PMC article.
-
Is the Philippines ready for the large-scale cultivation of Bt cotton?GM Crops Food. 2025 Dec;16(1):360-365. doi: 10.1080/21645698.2025.2499997. Epub 2025 May 2. GM Crops Food. 2025. PMID: 40316904 Free PMC article.
References
-
- Wendel J.F., Grover C.E. Taxonomy and Evolution of the Cotton Genus. In: Fang D.D., Percy R.G., editors. Gossypium. American Society of Agronomy, Inc.; Crop Science Society of America, Inc.; Soil Science Society of America, Inc.; Madison, WI, USA: 2015. pp. 25–44. Agronomy Monographs. (In Cotton) - DOI
LinkOut - more resources
Full Text Sources

