Structural Assembly of Qβ Virion and Its Diverse Forms of Virus-like Particles
- PMID: 35215818
- PMCID: PMC8880383
- DOI: 10.3390/v14020225
Structural Assembly of Qβ Virion and Its Diverse Forms of Virus-like Particles
Abstract
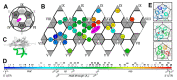
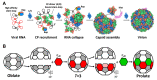
The coat proteins (CPs) of single-stranded RNA bacteriophages (ssRNA phages) directly assemble around the genomic RNA (gRNA) to form a near-icosahedral capsid with a single maturation protein (Mat) that binds the gRNA and interacts with the retractile pilus during infection of the host. Understanding the assembly of ssRNA phages is essential for their use in biotechnology, such as RNA protection and delivery. Here, we present the complete gRNA model of the ssRNA phage Qβ, revealing that the 3' untranslated region binds to the Mat and the 4127 nucleotides fold domain-by-domain, and is connected through long-range RNA-RNA interactions, such as kissing loops. Thirty-three operator-like RNA stem-loops are located and primarily interact with the asymmetric A/B CP-dimers, suggesting a pathway for the assembly of the virions. Additionally, we have discovered various forms of the virus-like particles (VLPs), including the canonical T = 3 icosahedral, larger T = 4 icosahedral, prolate, oblate forms, and a small prolate form elongated along the 3-fold axis. These particles are all produced during a normal infection, as well as when overexpressing the CPs. When overexpressing the shorter RNA fragments encoding only the CPs, we observed an increased percentage of the smaller VLPs, which may be sufficient to encapsidate a shorter RNA.
Keywords: genomic RNA; single-particle; ssRNA virus; virus assembly.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript or in the decision to publish the results.
Figures



Similar articles
-
Asymmetric cryo-EM structure of the canonical Allolevivirus Qβ reveals a single maturation protein and the genomic ssRNA in situ.Proc Natl Acad Sci U S A. 2016 Oct 11;113(41):11519-11524. doi: 10.1073/pnas.1609482113. Epub 2016 Sep 26. Proc Natl Acad Sci U S A. 2016. PMID: 27671640 Free PMC article.
-
In situ structures of the genome and genome-delivery apparatus in a single-stranded RNA virus.Nature. 2017 Jan 5;541(7635):112-116. doi: 10.1038/nature20589. Epub 2016 Dec 19. Nature. 2017. PMID: 27992877 Free PMC article.
-
Structures of Qβ virions, virus-like particles, and the Qβ-MurA complex reveal internal coat proteins and the mechanism of host lysis.Proc Natl Acad Sci U S A. 2017 Oct 31;114(44):11697-11702. doi: 10.1073/pnas.1707102114. Epub 2017 Oct 16. Proc Natl Acad Sci U S A. 2017. PMID: 29078304 Free PMC article.
-
Cryo-EM reveals infection steps of single-stranded RNA bacteriophages.Prog Biophys Mol Biol. 2021 Mar;160:79-86. doi: 10.1016/j.pbiomolbio.2020.07.011. Epub 2020 Aug 22. Prog Biophys Mol Biol. 2021. PMID: 32841651 Review.
-
Asymmetry in icosahedral viruses.Curr Opin Virol. 2019 Jun;36:67-73. doi: 10.1016/j.coviro.2019.05.006. Epub 2019 Jun 28. Curr Opin Virol. 2019. PMID: 31255982 Review.
Cited by
-
Evolutionary Qβ Phage Displayed Nanotag Library and Peptides for Biosensing.Viruses. 2023 Jun 22;15(7):1414. doi: 10.3390/v15071414. Viruses. 2023. PMID: 37515102 Free PMC article.
-
Heterologous Prime-Boost with Immunologically Orthogonal Protein Nanoparticles for Peptide Immunofocusing.bioRxiv [Preprint]. 2024 Feb 26:2024.02.24.581861. doi: 10.1101/2024.02.24.581861. bioRxiv. 2024. Update in: ACS Nano. 2024 Jul 23. doi: 10.1021/acsnano.4c00949. PMID: 38464232 Free PMC article. Updated. Preprint.
-
Coupling of Size Exclusion Chromatography to High Throughput Charge Detection Mass Spectrometry for the Analysis of Large Proteins and Virus-like Particles.Anal Chem. 2025 Feb 11;97(5):3036-3044. doi: 10.1021/acs.analchem.4c06084. Epub 2025 Jan 29. Anal Chem. 2025. PMID: 39878296 Free PMC article.
-
The diversity of protein-protein interaction interfaces within T=3 icosahedral viral capsids.Front Mol Biosci. 2022 Oct 20;9:967877. doi: 10.3389/fmolb.2022.967877. eCollection 2022. Front Mol Biosci. 2022. PMID: 36339706 Free PMC article.
-
A self-adjuvanted VLPs-based Covid-19 vaccine proven versatile, safe, and highly protective.Sci Rep. 2024 Oct 16;14(1):24228. doi: 10.1038/s41598-024-76163-w. Sci Rep. 2024. PMID: 39414952 Free PMC article.
References
-
- Van Duin J., Tsareva N. Single-stranded RNA phages. In: Calendar R., editor. The Bacteriophages. 2nd ed. Oxford University Press; New York, NY, USA: 2006. pp. 175–196.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

