Role of ubiquitin-protein ligase UBR5 in the disassembly of mitotic checkpoint complexes
- PMID: 35217622
- PMCID: PMC8892521
- DOI: 10.1073/pnas.2121478119
Role of ubiquitin-protein ligase UBR5 in the disassembly of mitotic checkpoint complexes
Abstract
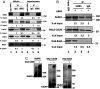
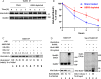
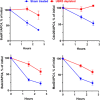
The mitotic (or spindle assembly) checkpoint system ensures accurate chromosome segregation in mitosis by preventing the onset of anaphase until correct bipolar attachment of sister chromosomes to the mitotic spindle is attained. It acts by promoting the assembly of a mitotic checkpoint complex (MCC), composed of mitotic checkpoint proteins BubR1, Bub3, Mad2, and Cdc20. MCC binds to and inhibits the action of ubiquitin ligase APC/C (anaphase-promoting complex/cyclosome), which targets for degradation regulators of anaphase initiation. When the checkpoint system is satisfied, MCCs are disassembled, allowing the recovery of APC/C activity and initiation of anaphase. Many of the pathways of the disassembly of the different MCCs have been elucidated, but the mode of their regulation remained unknown. We find that UBR5 (ubiquitin-protein ligase N-recognin 5) is associated with the APC/C*MCC complex immunopurified from extracts of nocodazole-arrested HeLa cells. UBR5 binds to mitotic checkpoint proteins BubR1, Bub3, and Cdc20 and promotes their polyubiquitylation in vitro. The dissociation of a Bub3*BubR1 subcomplex of MCC is stimulated by UBR5-dependent ubiquitylation, as suggested by observations that this process in mitotic extracts requires UBR5 and α-β bond hydrolysis of adenosine triphosphate. Furthermore, a system reconstituted from purified recombinant components carries out UBR5- and ubiquitylation-dependent dissociation of Bub3*BubR1. Immunodepletion of UBR5 from mitotic extracts slows down the release of MCC components from APC/C and prolongs the lag period in the recovery of APC/C activity in the exit from mitotic checkpoint arrest. We suggest that UBR5 may be involved in the regulation of the inactivation of the mitotic checkpoint.
Keywords: cell cycle; mitosis; ubiquitin.
Copyright © 2022 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures

References
-
- Musacchio A., Salmon E. D., The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8, 379–393 (2007). - PubMed
-
- Lara-Gonzalez P., Westhorpe F. G., Taylor S. S., The spindle assembly checkpoint. Curr. Biol. 22, R966–R980 (2012). - PubMed
-
- Musacchio A., The molecular biology of spindle assembly checkpoint signaling dynamics. Curr. Biol. 25, R1002–R1018 (2015). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

