Putting Humpty Dumpty Back Together Again: What Does Protein Quantification Mean in Bottom-Up Proteomics?
- PMID: 35220718
- PMCID: PMC8976764
- DOI: 10.1021/acs.jproteome.1c00894
Putting Humpty Dumpty Back Together Again: What Does Protein Quantification Mean in Bottom-Up Proteomics?
Abstract
Bottom-up proteomics provides peptide measurements and has been invaluable for moving proteomics into large-scale analyses. Commonly, a single quantitative value is reported for each protein-coding gene by aggregating peptide quantities into protein groups following protein inference or parsimony. However, given the complexity of both RNA splicing and post-translational protein modification, it is overly simplistic to assume that all peptides that map to a singular protein-coding gene will demonstrate the same quantitative response. By assuming that all peptides from a protein-coding sequence are representative of the same protein, we may miss the discovery of important biological differences. To capture the contributions of existing proteoforms, we need to reconsider the practice of aggregating protein values to a single quantity per protein-coding gene.
Keywords: post-translational modifications; protein grouping; proteoforms; quantitative analysis; quantitative proteomics.
Figures
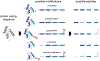
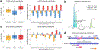
Similar articles
-
How can platelet proteomics best be used to interrogate disease?Platelets. 2023 Dec;34(1):2220046. doi: 10.1080/09537104.2023.2220046. Platelets. 2023. PMID: 37272536
-
Profiling proteoforms: promising follow-up of proteomics for biomarker discovery.Expert Rev Proteomics. 2014 Feb;11(1):121-9. doi: 10.1586/14789450.2014.878652. Expert Rev Proteomics. 2014. PMID: 24437377 Review.
-
Improving Proteoform Identifications in Complex Systems Through Integration of Bottom-Up and Top-Down Data.J Proteome Res. 2020 Aug 7;19(8):3510-3517. doi: 10.1021/acs.jproteome.0c00332. Epub 2020 Jul 10. J Proteome Res. 2020. PMID: 32584579 Free PMC article.
-
Bayesian proteoform modeling improves protein quantification of global proteomic measurements.Mol Cell Proteomics. 2014 Dec;13(12):3639-46. doi: 10.1074/mcp.M113.030932. Mol Cell Proteomics. 2014. PMID: 25433089 Free PMC article.
-
Proteomics beyond trypsin.FEBS J. 2015 Jul;282(14):2612-26. doi: 10.1111/febs.13287. Epub 2015 Apr 14. FEBS J. 2015. PMID: 25823410 Review.
Cited by
-
Inside the chase after those elusive proteoforms.Nat Methods. 2024 Feb;21(2):158-163. doi: 10.1038/s41592-024-02170-4. Nat Methods. 2024. PMID: 38308015 No abstract available.
-
Proteomics-The State of the Field: The Definition and Analysis of Proteomes Should Be Based in Reality, Not Convenience.Proteomes. 2024 Apr 19;12(2):14. doi: 10.3390/proteomes12020014. Proteomes. 2024. PMID: 38651373 Free PMC article.
-
scplainer: using linear models to understand mass spectrometry-based single-cell proteomics data.Genome Biol. 2025 Aug 7;26(1):237. doi: 10.1186/s13059-025-03713-4. Genome Biol. 2025. PMID: 40775377 Free PMC article.
-
Identification of Splice Variants and Isoforms in Transcriptomics and Proteomics.Annu Rev Biomed Data Sci. 2023 Aug 10;6:357-376. doi: 10.1146/annurev-biodatasci-020722-044021. Annu Rev Biomed Data Sci. 2023. PMID: 37561601 Free PMC article. Review.
-
Top-down proteomics of myosin light chain isoforms define chamber-specific expression in the human heart.J Mol Cell Cardiol. 2023 Aug;181:89-97. doi: 10.1016/j.yjmcc.2023.06.003. Epub 2023 Jun 14. J Mol Cell Cardiol. 2023. PMID: 37327991 Free PMC article.
References
-
- Nesvizhskii AI, Keller A, Kolker E & Aebersold R A Statistical Model for Identifying Proteins by Tandem Mass Spectrometry. Anal. Chem 75, 4646–4658 (2003). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

