A standardized assay for the quantitative detection of serum HBV RNA in chronic hepatitis B patients
- PMID: 35220917
- PMCID: PMC8920369
- DOI: 10.1080/22221751.2022.2045874
A standardized assay for the quantitative detection of serum HBV RNA in chronic hepatitis B patients
Abstract
Serum hepatitis B virus (HBV) pregenomic RNA (pgRNA) is a surrogate marker for reflecting the transcriptional activity of covalently closed circular DNA. However, there is still no standardized assay for the quantitative detection of serum HBV RNA in chronic hepatitis B patients. In this study, quantitative polymerase chain reactions for detecting the preC/C-RNA (preC/C region HBV pgRNA), SF-RNA (splicing variants-free pgRNA) and XR-RNA (X region remained pgRNA) regions were set up. The dynamic changes of serum pgRNA splicing variants and 3' terminal truncations were analysed in three retrospective cohorts: 35 treatment-naive chronic HBV-infected patients (cohort A), 52 chronic hepatitis B (CHB) patients who received nucleos(t)ide analogs (NAs) therapy for 48 weeks (cohort B) and eight CHB patients who are under long-term NAs treatment (cohort C). The accuracy and sensitivity of HBV RNA detection were assessed by the National Standard of HBV RNA. We confirmed that high proportions of pgRNA splicing variants and 3' terminal truncations were present and significantly affect the quantitative detection of serum HBV RNA in both treatment-naive and NAs-treated CHB patients. To achieve the higher accuracy and sensitivity on the detection of HBV RNA level, the primers and probes should be designed at the 5' terminal region of HBV genome and outside the mainly spliced sequence of pgRNA, especially for CHB patients under long-term NAs treatment. This study would help to better understand the significance of the pgRNA splicing variants and 3' terminal truncations, and further guide the clinical detection of serum HBV RNA.
Keywords: 3′ terminal truncations; HBV RNA; nucleos(t)ide analogs; quantitative detection; splicing variants.
Conflict of interest statement
One patent which belongs to Peking University of China is relevant to this work. GY, RC, YL, JZ, ZG, JW and FL are employees of Peking University.
Figures

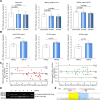


References
-
- Trépo C, Chan HL, Lok A.. Hepatitis B virus infection. Lancet (London, England). 2014 Dec 6;384(9959):2053–2063. - PubMed
-
- Wang J, Shen T, Huang X, et al. . Serum hepatitis B virus RNA is encapsidated pregenome RNA that may be associated with persistence of viral infection and rebound. J Hepatol. 2016 Oct;65(4):700–710. - PubMed
-
- Jansen L, Kootstra NA, van Dort KA, et al. . Hepatitis b virus pregenomic RNA Is present in Virions in plasma and is associated with a response to pegylated interferon alfa-2a and nucleos(t)ide analogues. J Infect Dis. 2016 Jan 15;213(2):224–232. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
