Cohesin-Mediated Chromatin Interactions and Autoimmunity
- PMID: 35222432
- PMCID: PMC8866859
- DOI: 10.3389/fimmu.2022.840002
Cohesin-Mediated Chromatin Interactions and Autoimmunity
Abstract
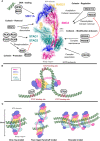
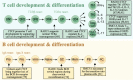
Proper physiological functioning of any cell type requires ordered chromatin organization. In this context, cohesin complex performs important functions preventing premature separation of sister chromatids after DNA replication. In partnership with CCCTC-binding factor, it ensures insulator activity to organize enhancers and promoters within regulatory chromatin. Homozygous mutations and dysfunction of individual cohesin proteins are embryonically lethal in humans and mice, which limits in vivo research work to embryonic stem cells and progenitors. Conditional alleles of cohesin complex proteins have been generated to investigate their functional roles in greater detail at later developmental stages. Thus, genome regulation enabled by action of cohesin proteins is potentially crucial in lineage cell development, including immune homeostasis. In this review, we provide current knowledge on the role of cohesin complex in leukocyte maturation and adaptive immunity. Conditional knockout and shRNA-mediated inhibition of individual cohesin proteins in mice demonstrated their importance in haematopoiesis, adipogenesis and inflammation. Notably, these effects occur rather through changes in transcriptional gene regulation than through expected cell cycle defects. This positions cohesin at the crossroad of immune pathways including NF-kB, IL-6, and IFNγ signaling. Cohesin proteins emerged as vital regulators at early developmental stages of thymocytes and B cells and after antigen challenge. Human genome-wide association studies are remarkably concordant with these findings and present associations between cohesin and rheumatoid arthritis, multiple sclerosis and HLA-B27 related chronic inflammatory conditions. Furthermore, bioinformatic prediction based on protein-protein interactions reveal a tight connection between the cohesin complex and immune relevant processes supporting the notion that cohesin will unearth new clues in regulation of autoimmunity.
Keywords: CCCTC-binding factor; autoimmunity; cohesin; genome organization; immune signaling.
Copyright © 2022 Chandrasekaran, Oparina, Garcia-Bonete, Wasén, Erlandsson, Malmhäll-Bah, Andersson, Jensen, Silfverswärd, Katona and Bokarewa.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

