Helicoverpa armigera miR-2055 regulates lipid metabolism via fatty acid synthase expression
- PMID: 35232249
- PMCID: PMC8889172
- DOI: 10.1098/rsob.210307
Helicoverpa armigera miR-2055 regulates lipid metabolism via fatty acid synthase expression
Abstract
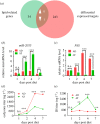
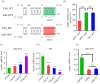
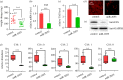
Insect hormones and microRNAs regulate lipid metabolism, but the mechanisms are not fully elucidated. Here, we found that cotton bollworm larvae feeding on Arabidopsis thaliana (AT) leaves had a lower triacylglycerol (TAG) level and more delayed development than individuals feeding on artificial diet (AD). Association analysis of small RNA and mRNA revealed that the level of miR-2055, a microRNA related to lipid metabolism, was significantly higher in larvae feeding on AT. Dual-luciferase reporter assays demonstrated miR-2055 binding to 3' UTR of fatty acid synthase (FAS) mRNA to suppress its expression. Elevating the level of miR-2055 in larvae by agomir injection decreased FAS mRNA and protein levels, which resulted in reduction of free fatty acid (FFA) and TAG in fat body. Interestingly, in vitro assays illustrated that juvenile hormone (JH) increased miR-2055 accumulation in a dosage-dependent manner, whereas knockdown of Methoprene tolerant (Met) or Kruppel homologue 1 (Kr-h1) decreased the miR-2055 level. This implied that JH induces the expression of miR-2055 via a Met-Kr-h1 signal. These findings demonstrate that JH and miRNA cooperate to modulate lipid synthesis, which provides new insights into the regulatory mechanisms of metabolism in insects.
Keywords: Helicoverpa armigera; development; juvenile hormone; lipid metabolism; microRNA.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
