SARS-CoV-2 ORF3a induces RETREG1/FAM134B-dependent reticulophagy and triggers sequential ER stress and inflammatory responses during SARS-CoV-2 infection
- PMID: 35239449
- PMCID: PMC9629087
- DOI: 10.1080/15548627.2022.2039992
SARS-CoV-2 ORF3a induces RETREG1/FAM134B-dependent reticulophagy and triggers sequential ER stress and inflammatory responses during SARS-CoV-2 infection
Abstract
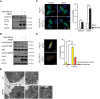
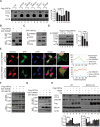
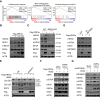
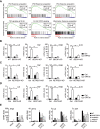
SARS-CoV-2 infections have resulted in a very large number of severe cases of COVID-19 and deaths worldwide. However, knowledge of SARS-CoV-2 infection, pathogenesis and therapy remains limited, emphasizing the urgent need for fundamental studies and drug development. Studies have shown that induction of macroautophagy/autophagy and hijacking of the autophagic machinery are essential for the infection and replication of SARS-CoV-2; however, the mechanism of this manipulation and the function of autophagy during SARS-CoV-2 infection remain unclear. In the present study, we identified ORF3a as an inducer of autophagy (in particular reticulophagy) and revealed that ORF3a localizes to the ER and induces RETREG1/FAM134B-related reticulophagy through the HMGB1-BECN1 (beclin 1) pathway. As a consequence, ORF3a induces ER stress and inflammatory responses through reticulophagy and then sensitizes cells to the acquisition of an ER stress-related early apoptotic phenotype and facilitates SARS-CoV-2 infection, suggesting that SARS-CoV-2 ORF3a hijacks reticulophagy and then disrupts ER homeostasis to induce ER stress and inflammatory responses during SARS-CoV-2 infection. These findings reveal the sequential induction of reticulophagy, ER stress and acute inflammatory responses during SARS-CoV-2 infection and imply the therapeutic potential of reticulophagy and ER stress-related drugs for COVID-19.Abbreviations: CQ: chloroquine; DEGs: differentially expressed genes; ER: endoplasmic reticulum; GSEA: gene set enrichment analysis; HMGB1: high mobility group box 1; HMOX1: heme oxygenase 1; MERS-CoV: Middle East respiratory syndrome coronavirus; RETREG1/FAM134B: reticulophagy regulator 1; RTN4: reticulon 4; SARS-CoV-2: severe acute respiratory syndrome coronavirus 2; TN: tunicamycin.
Keywords: ER stress; ORF3a; SARS-CoV-2; inflammatory response; reticulophagy.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures

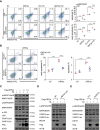
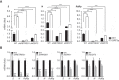

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous
