Genetic and hosts characterization of hantaviruses in port areas in Hainan Province, P. R. China
- PMID: 35239751
- PMCID: PMC8893628
- DOI: 10.1371/journal.pone.0264859
Genetic and hosts characterization of hantaviruses in port areas in Hainan Province, P. R. China
Abstract
Background: Hantaviruses (HVs) are major zoonotic pathogens in China that cause hemorrhagic fever with renal syndrome (HFRS) posing a major threat to people's health. Hainan Province, an island located in Southeast China, is an ideal region for sea ports. The unique tropical monsoon climate in Hainan provides sufficient living conditions for rodents, which help spread HVs and other rodent-borne diseases. In the routine monitoring of hantavirus, there was no evidence that rodents in Hainan carried hantavirus. No patients infected with hantavirus were found in the past. However, the surveillance of HVs-carrying rodents covering the whole territory of Hainan has not stopped.
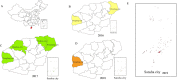
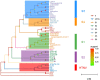
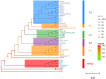
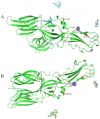
Methodology/principal findings: For the monitoring of the prevalence of HVs in rodents and the search for theoretical reference for rodent control and HFRS prevention, a total of 60 rodents from 6 monitoring spots were trapped around main ports in Hainan between 2016 and 2019. HV positive samples were identified by a specific kit and sequenced. The data indicated that seven rodents (Rattus norvegicus) were positive for hantavirus with a positivity rate of 11.67%. Phylogenetic analysis suggested that the two complete sequence strains HN1 and HN4 in this research were highly similar to the sequence strains GZRn36 and GZRn148 isolated in Guangdong Province, and they located in the same phylogenetic tree branch which belongs to S2 subtype. Although the two partial sequences HT1 and HT2 isolated in Xisha Islands belong to S2 subtype according to the phylogenetic tree of L segment, they showed a great nucleotide difference with HN1 and HN4. We also found 13 amino acid variations compared with SEOV 80-39 and 6 amino acid mutations related to epitope, and the variations may reduce the effectiveness of the current HFRS vaccines used in humans.
Conclusions/significance: The study indicated HVs carried by rodents found in Hainan Province may be transmitted from Guangdong Province through trading ports and carriage of goods by sea. So it is of great significance to strengthen the surveillance of rodents in port areas especially capture and eliminate rodents on ship. Timely elimination of host animals of hantavirus in port areas is necessary to prevent an outbreak of HVs disease.
Conflict of interest statement
All the authors declared no competing interests.
Figures


References
-
- Wang Q, Yue M, Yao P, Zhu C, Ai L, Hu D, et al. Epidemic Trend and Molecular Evolution of HV Family in the Main Hantavirus Epidemic Areas From 2004 to 2016, in P.R. China. Front Cell Infect Microbiol. 2020;10:584814. Epub 2021/02/23. doi: 10.3389/fcimb.2020.584814 ; PubMed Central PMCID: PMC7886990. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

