Comparison between short-term stress and long-term adaptive responses reveal common paths to molecular adaptation
- PMID: 35243257
- PMCID: PMC8873613
- DOI: 10.1016/j.isci.2022.103899
Comparison between short-term stress and long-term adaptive responses reveal common paths to molecular adaptation
Abstract
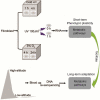
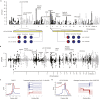
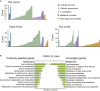
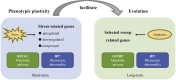
The phenotypic plasticity in responses to short-term stress can provide clues for understanding the adaptive fixation mechanism of genetic variation during long-term exposure to extreme environments. However, few studies have compared short-term stress responses with long-term evolutionary patterns; in particular, no interactions between the two processes have been evaluated in high-altitude environment. We performed RNA sequencing in embryo fibroblasts derived from great tits and mice to explore transcriptional responses after exposure to simulated high-altitude environmental stresses. Transcriptional changes of genes associated with metabolic pathways were identified in both bird and mice cells after short-term stress responses. Genomic comparisons among long-term highland tits and mammals and their lowland relatives revealed similar pathways (e.g., metabolic pathways) with that initiated under short-term stress transcriptional responses in vitro. These findings highlight the indicative roles of short-term stress in the long-term adaptation, and adopt common paths to molecular adaptation in mouse and bird cells.
Keywords: Biological sciences; Cell biology; Genetics; Genomics.
© 2022 The Authors.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Functional genomics of adaptation to hypoxic cold-stress in high-altitude deer mice: transcriptomic plasticity and thermogenic performance.Evolution. 2014 Jan;68(1):48-62. doi: 10.1111/evo.12257. Epub 2013 Sep 16. Evolution. 2014. PMID: 24102503 Free PMC article.
-
Physiological and genomic evidence that selection on the transcription factor Epas1 has altered cardiovascular function in high-altitude deer mice.PLoS Genet. 2019 Nov 7;15(11):e1008420. doi: 10.1371/journal.pgen.1008420. eCollection 2019 Nov. PLoS Genet. 2019. PMID: 31697676 Free PMC article.
-
Transcriptomes of Saussurea (Asteraceae) Provide Insights into High-Altitude Adaptation.Plants (Basel). 2021 Aug 20;10(8):1715. doi: 10.3390/plants10081715. Plants (Basel). 2021. PMID: 34451759 Free PMC article.
-
Human Genetic Adaptation to High Altitude: Evidence from the Andes.Genes (Basel). 2019 Feb 15;10(2):150. doi: 10.3390/genes10020150. Genes (Basel). 2019. PMID: 30781443 Free PMC article. Review.
-
Functional Genomic Insights into Regulatory Mechanisms of High-Altitude Adaptation.Adv Exp Med Biol. 2016;903:113-28. doi: 10.1007/978-1-4899-7678-9_8. Adv Exp Med Biol. 2016. PMID: 27343092 Free PMC article. Review.
Cited by
-
A time-resolved multi-omics atlas of transcriptional regulation in response to high-altitude hypoxia across whole-body tissues.Nat Commun. 2024 May 10;15(1):3970. doi: 10.1038/s41467-024-48261-w. Nat Commun. 2024. PMID: 38730227 Free PMC article.
-
The metabolic adaptation in wild vertebrates via omics approaches.Life Metab. 2022 Dec 28;1(3):234-241. doi: 10.1093/lifemeta/loac040. eCollection 2022 Dec. Life Metab. 2022. PMID: 39872075 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

