Downregulation of ATP binding cassette subfamily a member 10 acts as a prognostic factor associated with immune infiltration in breast cancer
- PMID: 35247251
- PMCID: PMC8954971
- DOI: 10.18632/aging.203933
Downregulation of ATP binding cassette subfamily a member 10 acts as a prognostic factor associated with immune infiltration in breast cancer
Abstract
The human ATP binding cassette (ABC) family of transporter proteins plays an important role in the maintenance of homeostasis in vivo. The aim of this study is to evaluate the potential diagnostic, prognostic, and therapeutic value of the ABCA10 gene in BRCA. We found that ABCA10 expression was downregulated in different subgroups of breast cancer and strongly correlated with pathological stage in BRCA patients. Low expression of ABCA10 was associated with BRCA patients showing shorter overall survival (OS). ABCA10 expression may be regulated by promoter methylation, copy number variation (CNV) and kinase, and is associated with immune infiltration. Our study also demonstrated the potential role of ABCA10 modifications in tumor microenvironment (TME) cellular infiltration. Nevertheless, the regulatory mechanism remains unknown and immunotherapy is marginal in BRCA. We demonstrate the expression of different ABCA10 modulators in breast cancer associated with genetic variants, deletions, tumor mutation burden (TMB) and TME. Mutations in ABCA10 are positively associated with different immune cells in six different immune databases and play an important role in immune cell infiltration in breast cancer. Overall, this study provides evidence that ABCA10 could become the potential targets for precision treatment and new biomarkers in the prognosis of breast cancer.
Keywords: ABCA10; biomarker; breast cancer; multi-omics; prognosis.
Conflict of interest statement
Figures
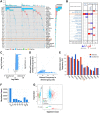









Similar articles
-
Integrated pan-cancer analysis revealed therapeutic targets in the ABC transporter protein family.PLoS One. 2025 May 30;20(5):e0308585. doi: 10.1371/journal.pone.0308585. eCollection 2025. PLoS One. 2025. PMID: 40445912 Free PMC article.
-
NEFM DNA methylation correlates with immune infiltration and survival in breast cancer.Clin Epigenetics. 2021 May 17;13(1):112. doi: 10.1186/s13148-021-01096-4. Clin Epigenetics. 2021. PMID: 34001208 Free PMC article.
-
ATP2C2 Has Potential to Define Tumor Microenvironment in Breast Cancer.Front Immunol. 2021 Apr 14;12:657950. doi: 10.3389/fimmu.2021.657950. eCollection 2021. Front Immunol. 2021. PMID: 33936088 Free PMC article.
-
Identification of Five Immune-Related lncRNAs Predicting Survival and Tumor Microenvironment Characteristics in Breast Cancer.Comput Math Methods Med. 2021 Feb 27;2021:6676692. doi: 10.1155/2021/6676692. eCollection 2021. Comput Math Methods Med. 2021. PMID: 33727952 Free PMC article.
-
The role of IL-35 and IL-37 in breast cancer - potential therapeutic targets for precision medicine.Front Oncol. 2022 Nov 22;12:1051282. doi: 10.3389/fonc.2022.1051282. eCollection 2022. Front Oncol. 2022. PMID: 36483045 Free PMC article. Review.
Cited by
-
Multi-omics and experimental analysis unveil theragnostic value and immunological roles of inner membrane mitochondrial protein (IMMT) in breast cancer.J Transl Med. 2023 Mar 10;21(1):189. doi: 10.1186/s12967-023-04035-4. J Transl Med. 2023. PMID: 36899366 Free PMC article.
-
Tumor-derived exosomes RNA expression profiling identifies the prognosis, immune characteristics, and treatment in HR+/HER2-breast cancer.Aging (Albany NY). 2023 Aug 29;15(16):8471-8486. doi: 10.18632/aging.204986. Epub 2023 Aug 29. Aging (Albany NY). 2023. PMID: 37647033 Free PMC article.
-
Mitochondrial calcium uniporter as biomarker and therapeutic target for breast cancer: Prognostication, immune microenvironment, epigenetic regulation and precision medicine.J Adv Res. 2025 Apr;70:445-461. doi: 10.1016/j.jare.2024.04.015. Epub 2024 Apr 24. J Adv Res. 2025. PMID: 38663838 Free PMC article.
-
Spatial and single-cell explorations uncover prognostic significance and immunological functions of mitochondrial calcium uniporter in breast cancer.Cancer Cell Int. 2024 Apr 17;24(1):140. doi: 10.1186/s12935-024-03327-z. Cancer Cell Int. 2024. PMID: 38632642 Free PMC article.
-
Expression profiling of luminal B breast tumor in Indian women.J Cancer Res Clin Oncol. 2023 Nov;149(15):13645-13664. doi: 10.1007/s00432-023-05195-y. Epub 2023 Jul 30. J Cancer Res Clin Oncol. 2023. PMID: 37516983 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

