Cardiac Dysfunction Promotes Cancer Progression via Multiple Secreted Factors
- PMID: 35260887
- PMCID: PMC9359722
- DOI: 10.1158/0008-5472.CAN-21-2463
Cardiac Dysfunction Promotes Cancer Progression via Multiple Secreted Factors
Abstract
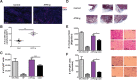
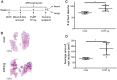
Heart failure and cancer are the leading cause of deaths worldwide. While heart failure and cancer have been considered separate diseases, it is becoming evident that they are highly connected and affect each other's outcomes. Recent studies using experimental mouse models have suggested that heart failure promotes tumor progression. The mouse models used involve major irreversible surgery. Here, we induced heart hypertrophy via expression of activating transcription factor 3 (ATF3) in cardiomyocytes, followed by cancer cells' implantation. Tumors developing in ATF3-transgenic mice grew larger and displayed a more highly metastatic phenotype compared with tumors in wild-type mice. To address whether ATF3 expression or the cardiac outcome are necessary for tumor progression, ATF3 expression was turned off after cardiac hypertrophy development followed by cancer cell implantation. The tumor promotion phenotype and the enhancement of metastatic properties were preserved, suggesting that the failing heart per se is sufficient to promote tumor progression. Serum derived from ATF3-transgenic mice enhanced cancer cell proliferation and increased cancer cell metastatic properties in vitro. Using a cytokine array panel, multiple factors responsible for promoting tumor cell proliferation and the metastatic phenotype were identified. Interestingly, the failing heart and the tumor separately and simultaneously contributed to higher levels of these factors in the serum as well as other tissues and organs. These data suggest the existence of intimate cross-talk between the hypertrophied heart and the tumor that is mediated by secreted factors, leading to cancer promotion and disease deterioration.
Significance: This work highlights the importance of early diagnosis and treatment of heart failure prior to reaching the irreversible stage that can exacerbate cancer progression.
©2022 The Authors; Published by the American Association for Cancer Research.
Figures



Similar articles
-
Induction of activating transcription factor 3 limits survival following infarct-induced heart failure in mice.Am J Physiol Heart Circ Physiol. 2015 Oct;309(8):H1326-35. doi: 10.1152/ajpheart.00513.2015. Epub 2015 Sep 4. Am J Physiol Heart Circ Physiol. 2015. PMID: 26342068 Free PMC article.
-
Adult cardiac expression of the activating transcription factor 3, ATF3, promotes ventricular hypertrophy.PLoS One. 2013 Jul 3;8(7):e68396. doi: 10.1371/journal.pone.0068396. Print 2013. PLoS One. 2013. PMID: 23874609 Free PMC article.
-
The cardiac maladaptive ATF3-dependent cross-talk between cardiomyocytes and macrophages is mediated by the IFNγ-CXCL10-CXCR3 axis.Int J Cardiol. 2017 Feb 1;228:394-400. doi: 10.1016/j.ijcard.2016.11.159. Epub 2016 Nov 10. Int J Cardiol. 2017. PMID: 27870968
-
ATF3-dependent cross-talk between cardiomyocytes and macrophages promotes cardiac maladaptive remodeling.Int J Cardiol. 2015 Nov 1;198:232-40. doi: 10.1016/j.ijcard.2015.06.099. Epub 2015 Jul 9. Int J Cardiol. 2015. PMID: 26201690
-
Molecular switches under TGFβ signalling during progression from cardiac hypertrophy to heart failure.Br J Pharmacol. 2016 Jan;173(1):3-14. doi: 10.1111/bph.13344. Epub 2015 Nov 16. Br J Pharmacol. 2016. PMID: 26431212 Free PMC article. Review.
Cited by
-
Cancer Development and Progression in Patients with Heart Failure.Curr Heart Fail Rep. 2024 Dec;21(6):515-529. doi: 10.1007/s11897-024-00680-y. Epub 2024 Sep 28. Curr Heart Fail Rep. 2024. PMID: 39340596 Free PMC article. Review.
-
The crosstalk of breast cancer and ischemic heart disease.Cell Death Discov. 2025 Apr 18;11(1):185. doi: 10.1038/s41420-025-02428-6. Cell Death Discov. 2025. PMID: 40251177 Free PMC article. Review.
-
Tumor Growth Ameliorates Cardiac Dysfunction.Cells. 2023 Jul 14;12(14):1853. doi: 10.3390/cells12141853. Cells. 2023. PMID: 37508517 Free PMC article.
-
Current evidence regarding the cellular mechanisms associated with cancer progression due to cardiovascular diseases.J Transl Med. 2024 Jan 26;22(1):105. doi: 10.1186/s12967-023-04803-2. J Transl Med. 2024. PMID: 38279150 Free PMC article. Review.
-
Common biological processes and mutual crosstalk mechanisms between cardiovascular disease and cancer.Front Oncol. 2024 Nov 20;14:1453090. doi: 10.3389/fonc.2024.1453090. eCollection 2024. Front Oncol. 2024. PMID: 39634266 Free PMC article. Review.
References
-
- Ponikowski P, Anker SD, AlHabib KF, Cowie MR, Force TL, Hu S, et al. . Heart failure: preventing disease and death worldwide. ESC Heart Fail 2014;1:4–25. - PubMed
-
- Tocchetti CG, Ameri P, de Boer RA, D'Alessandra Y, Russo M, Sorriento D, et al. . Cardiac dysfunction in cancer patients: beyond direct cardiomyocyte damage of anticancer drugs: novel cardio-oncology insights from the joint 2019 meeting of the ESC Working Groups of Myocardial Function and Cellular Biology of the Heart. Cardiovasc Res 2020;116:1820–34. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

