Approach to map nanotopography of cell surface receptors
- PMID: 35264712
- PMCID: PMC8907216
- DOI: 10.1038/s42003-022-03152-y
Approach to map nanotopography of cell surface receptors
Abstract
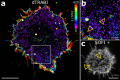
Cells communicate with their environment via surface receptors, but nanoscopic receptor organization with respect to complex cell surface morphology remains unclear. This is mainly due to a lack of accessible, robust and high-resolution methods. Here, we present an approach for mapping the topography of receptors at the cell surface with nanometer precision. The method involves coating glass coverslips with glycine, which preserves the fine membrane morphology while allowing immobilized cells to be positioned close to the optical surface. We developed an advanced and simplified algorithm for the analysis of single-molecule localization data acquired in a biplane detection scheme. These advancements enable direct and quantitative mapping of protein distribution on ruffled plasma membranes with near isotropic 3D nanometer resolution. As demonstrated successfully for CD4 and CD45 receptors, the described workflow is a straightforward quantitative technique to study molecules and their interactions at the complex surface nanomorphology of differentiated metazoan cells.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
Single-molecule imaging of cell surfaces using near-field nanoscopy.Acc Chem Res. 2012 Mar 20;45(3):327-36. doi: 10.1021/ar2001167. Epub 2011 Oct 12. Acc Chem Res. 2012. PMID: 21992025 Review.
-
Nanoscale organization of the pathogen receptor DC-SIGN mapped by single-molecule high-resolution fluorescence microscopy.Chemphyschem. 2007 Jul 16;8(10):1473-80. doi: 10.1002/cphc.200700169. Chemphyschem. 2007. PMID: 17577901
-
Mapping molecular diffusion in the plasma membrane by Multiple-Target Tracing (MTT).J Vis Exp. 2012 May 27;(63):e3599. doi: 10.3791/3599. J Vis Exp. 2012. PMID: 22664619 Free PMC article.
-
Mapping the Nicotinic Acetylcholine Receptor Nanocluster Topography at the Cell Membrane with STED and STORM Nanoscopies.Int J Mol Sci. 2022 Sep 9;23(18):10435. doi: 10.3390/ijms231810435. Int J Mol Sci. 2022. PMID: 36142349 Free PMC article.
-
Micro-nanopatterning as tool to study the role of physicochemical properties on cell-surface interactions.J Biomed Mater Res A. 2013 Oct;101(10):3019-32. doi: 10.1002/jbm.a.34586. Epub 2013 Apr 5. J Biomed Mater Res A. 2013. PMID: 23559501 Review.
Cited by
-
Cryosectioning-enhanced super-resolution microscopy for single-protein imaging across cells and tissues.bioRxiv [Preprint]. 2025 Feb 13:2024.02.05.576943. doi: 10.1101/2024.02.05.576943. bioRxiv. 2025. Update in: Proc Natl Acad Sci U S A. 2025 Aug 12;122(32):e2504578122. doi: 10.1073/pnas.2504578122. PMID: 38370628 Free PMC article. Updated. Preprint.
-
Cryosectioning-enhanced super-resolution microscopy for single-protein imaging across cells and tissues.Proc Natl Acad Sci U S A. 2025 Aug 12;122(32):e2504578122. doi: 10.1073/pnas.2504578122. Epub 2025 Aug 7. Proc Natl Acad Sci U S A. 2025. PMID: 40773232 Free PMC article.
-
Plasma membrane topography governs the 3D dynamic localization of IgM B cell antigen receptor clusters.EMBO J. 2023 Feb 15;42(4):e112030. doi: 10.15252/embj.2022112030. Epub 2023 Jan 3. EMBO J. 2023. PMID: 36594262 Free PMC article.
-
Role of Lipids in Morphogenesis of T-Cell Microvilli.Front Immunol. 2021 Mar 10;12:613591. doi: 10.3389/fimmu.2021.613591. eCollection 2021. Front Immunol. 2021. PMID: 33790891 Free PMC article. Review.
-
Cell surface morphology mimicking nano-bio platform for immune cell stimulation.iScience. 2024 Sep 26;27(11):111033. doi: 10.1016/j.isci.2024.111033. eCollection 2024 Nov 15. iScience. 2024. PMID: 39498306 Free PMC article.
References
-
- Cebecauer M, Spitaler M, Serge A, Magee AI. Signalling complexes and clusters: functional advantages and methodological hurdles. J. Cell Sci. 2010;123:309–320. - PubMed
-
- Rossier O, et al. Integrins beta1 and beta3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat. Cell Biol. 2012;14:1057–1067. - PubMed
-
- Kellermayer B, et al. Differential nanoscale topography and functional role of GluN2-NMDA receptor subtypes at glutamatergic synapses. Neuron. 2018;100:106–119 e7. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- SBF003\1163/Wellcome Trust (Wellcome)
- WT_/Wellcome Trust/United Kingdom
- 19-0704S/Grantová Agentura České Republiky (Grant Agency of the Czech Republic)
- EP/N509760/1/RCUK | Engineering and Physical Sciences Research Council (EPSRC)
- 2031229/RCUK | Engineering and Physical Sciences Research Council (EPSRC)
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

