Faecal microbiome and metabolic signatures in rectal neuroendocrine tumors
- PMID: 35265196
- PMCID: PMC8899573
- DOI: 10.7150/thno.66464
Faecal microbiome and metabolic signatures in rectal neuroendocrine tumors
Abstract
Background: The prevalence of rectal neuroendocrine tumors (RNET) has increased substantially over the past decades. Little is known on mechanistic alteration in the pathogenesis of such disease. We postulate that perturbations of human gut microbiome-metabolome interface influentially affect the development of RNET. The study aims to characterize the composition and function of faecal microbiome and metabolites in RNET individuals. Methods: We performed deep shotgun metagenomic sequencing and untargeted liquid chromatography-mass spectrometry (LC-MS) metabolomic profiling of faecal samples from the discovery cohort (18 RNET patients, 40 controls), and validated the microbiome and metabolite-based classifiers in an independent cohort (15 RNET participants, 19 controls). Results: We uncovered a dysbiotic gut ecological microenvironment in RNET patients, characterized by aberrant depletion and attenuated connection of microbial species, and abnormally aggregated lipids and lipid-like molecules. Functional characterization based on our in-house and Human Project Unified Metabolic Analysis Network 2 (HUMAnN2) pipelines further indicated a nutrient deficient gut microenvironment in RNET individuals, evidenced by diminished activities such as energy metabolism, vitamin biosynthesis and transportation. By integrating these data, we revealed 291 robust associations between representative differentially abundant taxonomic species and metabolites, indicating a tight interaction of gut microbiome with metabolites in RNET pathogenesis. Finally, we identified a cluster of gut microbiome and metabolite-based signatures, and replicated them in an independent cohort, showing accurate prediction of such neoplasm from healthy people. Conclusions: Our current study is the first to comprehensively characterize the perturbed interface of gut microbiome and metabolites in RNET patients, which may provide promising targets for microbiome-based diagnostics and therapies for this disorder.
Keywords: Gut microbiota; Metabolite; Rectal neuroendocrine tumor; Shotgun metagenomic sequencing; Untargeted liquid chromatography-mass spectrometry (LC-MS) metabolomics..
© The author(s).
Conflict of interest statement
Competing Interests: The authors have declared that no competing interest exists.
Figures



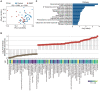


References
-
- Patel N, Barbieri A, Gibson J. Neuroendocrine Tumors of the Gastrointestinal Tract and Pancreas. Surg Pathol Clin. 2019;12:1021–44. - PubMed
-
- Fraenkel M, Kim M, Faggiano A, de Herder WW, Valk GD, Knowledge N. Incidence of gastroenteropancreatic neuroendocrine tumours: a systematic review of the literature. Endocr Relat Cancer. 2014;21:R153–63. - PubMed
-
- Cives M, Strosberg JR. Gastroenteropancreatic Neuroendocrine Tumors. CA Cancer J Clin. 2018;68:471–87. - PubMed
-
- Starzynska T, Londzin-Olesik M, Baldys-Waligorska A, Bednarczuk T, Blicharz-Dorniak J, Bolanowski M. et al. Colorectal neuroendocrine neoplasms - management guidelines (recommended by the Polish Network of Neuroendocrine Tumours) Endokrynol Pol. 2017;68:250–60. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

