Diversity of bet-hedging strategies in microbial communities-Recent cases and insights
- PMID: 35266649
- PMCID: PMC9286555
- DOI: 10.1002/wsbm.1544
Diversity of bet-hedging strategies in microbial communities-Recent cases and insights
Abstract
Microbial communities are continuously exposed to unpredictable changes in their environment. To thrive in such dynamic habitats, microorganisms have developed the ability to readily switch phenotypes, resulting in a number of differently adapted subpopulations expressing various traits. In evolutionary biology, a particular case of phenotypic heterogeneity that evolved in an unpredictably changing environment has been defined as bet-hedging. Bet-hedging is a risk-spreading strategy where isogenic populations stochastically (randomly) diversify their phenotypes, often resulting in maladapted individuals that suffer lower reproductive success. This fitness trade-off in a specific environment may have a selective advantage upon the sudden environmental shift. Thus, a bet-hedging strategy allows populations to persist in very dynamic habitats, but with a particular fitness cost. In recent years, numerous examples of phenotypic heterogeneity in different microorganisms have been observed, some suggesting bet-hedging. Here, we highlight the latest reports concerning bet-hedging phenomena in various microorganisms to show how versatile this strategy is within the microbial realms. This article is categorized under: Infectious Diseases > Molecular and Cellular Physiology.
Keywords: adaptation; bet-hedging; evolutionary strategy; persisters; phenotypic heterogeneity.
© 2021 The Authors. WIREs Mechanisms of Disease published by Wiley Periodicals LLC.
Conflict of interest statement
The authors have declared no conflicts of interest for this article.
Figures



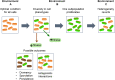
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

