Development of Cas12a-Based Cell-Free Small-Molecule Biosensors via Allosteric Regulation of CRISPR Array Expression
- PMID: 35266687
- PMCID: PMC8943526
- DOI: 10.1021/acs.analchem.1c04332
Development of Cas12a-Based Cell-Free Small-Molecule Biosensors via Allosteric Regulation of CRISPR Array Expression
Abstract
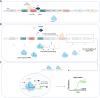
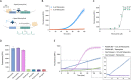
Cell-free biosensors can detect various molecules, thus promising to transform the landscape of diagnostics. Here, we developed a simple, rapid, sensitive, and field-deployable small-molecule detection platform based on allosteric transcription factor (aTF)-regulated expression of a clustered regularly interspaced short palindromic repeats (CRISPR) array coupled to Cas12a activity. To this end, we engineered an expression cassette harboring a T7 promoter, an aTF binding sequence, a Cas12a CRISPR array, and protospacer adjacent motif-flanked Cas12a target sequences. In the presence of the ligand, dissociation of the aTF allows transcription of the CRISPR array; this leads to activation of Cas12a collateral activity, which cleaves a single-stranded DNA linker to free a quenched fluorophore, resulting in a rapid, significant increase of fluorescence. As a proof of concept, we used TetR as the aTF to detect different tetracycline antibiotics with high sensitivity and specificity and a simple, hand-held visualizer to develop a fluorescence-based visual readout. We also adapted a mobile phone application to further simplify the interpretation of the results. Finally, we showed that the reagents could be lyophilized to facilitate storage and distribution. This detection platform represents a valuable addition to the toolbox of cell-free, CRISPR-based biosensors, with great potential for in-field deployment to detect non-nucleic acid small molecules.
Conflict of interest statement
The authors declare no competing financial interest.
Figures


Similar articles
-
Aptamer assisted CRISPR-Cas12a strategy for small molecule diagnostics.Biosens Bioelectron. 2021 Jul 1;183:113196. doi: 10.1016/j.bios.2021.113196. Epub 2021 Mar 24. Biosens Bioelectron. 2021. PMID: 33839534
-
CRISPR-Cas12a System for Biosensing and Gene Regulation.Chem Asian J. 2021 Apr 19;16(8):857-867. doi: 10.1002/asia.202100043. Epub 2021 Mar 18. Chem Asian J. 2021. PMID: 33638271 Review.
-
Detection of Cancer Marker Flap Endonuclease 1 Using One-Pot Transcription-Powered Clustered Regularly Interspaced Short Palindromic Repeat/Cas12a Signal Expansion.Anal Chem. 2022 Oct 4;94(39):13549-13555. doi: 10.1021/acs.analchem.2c03054. Epub 2022 Sep 19. Anal Chem. 2022. PMID: 36121799
-
G-triplex: A new type of CRISPR-Cas12a reporter enabling highly sensitive nucleic acid detection.Biosens Bioelectron. 2021 Sep 1;187:113292. doi: 10.1016/j.bios.2021.113292. Epub 2021 May 9. Biosens Bioelectron. 2021. PMID: 33991961
-
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas Advancement in Molecular Diagnostics and Signal Readout Approaches.J Mol Diagn. 2021 Nov;23(11):1433-1442. doi: 10.1016/j.jmoldx.2021.07.025. Epub 2021 Aug 25. J Mol Diagn. 2021. PMID: 34454111 Review.
Cited by
-
CRISPR Assays for Disease Diagnosis: Progress to and Barriers Remaining for Clinical Applications.Adv Sci (Weinh). 2023 Jul;10(20):e2301697. doi: 10.1002/advs.202301697. Epub 2023 May 10. Adv Sci (Weinh). 2023. PMID: 37162202 Free PMC article. Review.
-
Bio-SCAN V2: A CRISPR/dCas9-based lateral flow assay for rapid detection of theophylline.Front Bioeng Biotechnol. 2023 Jan 19;11:1118684. doi: 10.3389/fbioe.2023.1118684. eCollection 2023. Front Bioeng Biotechnol. 2023. PMID: 36741753 Free PMC article.
-
Advancements and Prospects in DNA-Based Bioanalytical Technology for Environmental Toxicant Detection.JACS Au. 2025 Jun 2;5(6):2443-2462. doi: 10.1021/jacsau.5c00398. eCollection 2025 Jun 23. JACS Au. 2025. PMID: 40575295 Free PMC article. Review.
-
Modern Plant Breeding Techniques in Crop Improvement and Genetic Diversity: From Molecular Markers and Gene Editing to Artificial Intelligence-A Critical Review.Plants (Basel). 2024 Sep 24;13(19):2676. doi: 10.3390/plants13192676. Plants (Basel). 2024. PMID: 39409546 Free PMC article. Review.
-
The CRISPR/Cas System: A Customizable Toolbox for Molecular Detection.Genes (Basel). 2023 Mar 31;14(4):850. doi: 10.3390/genes14040850. Genes (Basel). 2023. PMID: 37107608 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

