Chromatin Remodeler Smarca5 Is Required for Cancer-Related Processes of Primary Cell Fitness and Immortalization
- PMID: 35269430
- PMCID: PMC8909548
- DOI: 10.3390/cells11050808
Chromatin Remodeler Smarca5 Is Required for Cancer-Related Processes of Primary Cell Fitness and Immortalization
Abstract
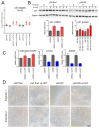
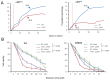
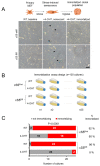
Smarca5, an ATPase of the ISWI class of chromatin remodelers, is a key regulator of chromatin structure, cell cycle and DNA repair. Smarca5 is deregulated in leukemia and breast, lung and gastric cancers. However, its role in oncogenesis is not well understood. Chromatin remodelers often play dosage-dependent roles in cancer. We therefore investigated the epigenomic and phenotypic impact of controlled stepwise attenuation of Smarca5 function in the context of primary cell transformation, a process relevant to tumor formation. Upon conditional single- or double-allele Smarca5 deletion, the cells underwent both accelerated growth arrest and senescence entry and displayed gradually increased sensitivity to genotoxic insults. These phenotypic characteristics were explained by specific remodeling of the chromatin structure and the transcriptome in primary cells prior to the immortalization onset. These molecular programs implicated Smarca5 requirement in DNA damage repair, telomere maintenance, cell cycle progression and in restricting apoptosis and cellular senescence. Consistent with the molecular programs, we demonstrate for the first time that Smarca5-deficient primary cells exhibit dramatically decreased capacity to bypass senescence and immortalize, an indispensable step during cell transformation and cancer development. Thus, Smarca5 plays a crucial role in key homeostatic processes and sustains cancer-promoting molecular programs and cellular phenotypes.
Keywords: ATAC-seq; MEF; RNA-seq; Smarca5; Snf2h; cell cycle; cell immortalization; homologous recombination; non-homologous end-joining; senescence.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Poly(ADP-ribosyl)ation links the chromatin remodeler SMARCA5/SNF2H to RNF168-dependent DNA damage signaling.J Cell Sci. 2013 Feb 15;126(Pt 4):889-903. doi: 10.1242/jcs.109413. Epub 2012 Dec 21. J Cell Sci. 2013. PMID: 23264744
-
NuMA promotes homologous recombination repair by regulating the accumulation of the ISWI ATPase SNF2h at DNA breaks.Nucleic Acids Res. 2014 Jun;42(10):6365-79. doi: 10.1093/nar/gku296. Epub 2014 Apr 20. Nucleic Acids Res. 2014. PMID: 24753406 Free PMC article.
-
Human ISWI complexes are targeted by SMARCA5 ATPase and SLIDE domains to help resolve lesion-stalled transcription.Nucleic Acids Res. 2014 Jul;42(13):8473-85. doi: 10.1093/nar/gku565. Epub 2014 Jul 2. Nucleic Acids Res. 2014. PMID: 24990377 Free PMC article.
-
An insight into understanding the coupling between homologous recombination mediated DNA repair and chromatin remodeling mechanisms in plant genome: an update.Cell Cycle. 2021 Sep;20(18):1760-1784. doi: 10.1080/15384101.2021.1966584. Epub 2021 Aug 26. Cell Cycle. 2021. PMID: 34437813 Free PMC article. Review.
-
Chromatin modification and NBS1: their relationship in DNA double-strand break repair.Genes Genet Syst. 2016;90(4):195-208. doi: 10.1266/ggs.15-00010. Epub 2015 Nov 25. Genes Genet Syst. 2016. PMID: 26616756 Review.
Cited by
-
Epigenetic modulators provide a path to understanding disease and therapeutic opportunity.Genes Dev. 2024 Jul 19;38(11-12):473-503. doi: 10.1101/gad.351444.123. Genes Dev. 2024. PMID: 38914477 Free PMC article. Review.
-
Hyperglycemia-Suppressed SMARCA5 Disrupts Transcriptional Homeostasis to Facilitate Endothelial Dysfunction in Diabetes.Diabetes Metab J. 2023 May;47(3):366-381. doi: 10.4093/dmj.2022.0179. Epub 2023 Mar 6. Diabetes Metab J. 2023. PMID: 36872061 Free PMC article.
-
Molecular mechanisms of circular RNA in breast cancer: a narrative review.Transl Cancer Res. 2024 Feb 29;13(2):1139-1149. doi: 10.21037/tcr-23-1760. Epub 2024 Feb 26. Transl Cancer Res. 2024. PMID: 38482408 Free PMC article. Review.
-
USP3 promotes DNA damage response and chemotherapy resistance through stabilizing and deubiquitinating SMARCA5 in prostate cancer.Cell Death Dis. 2024 Nov 5;15(11):790. doi: 10.1038/s41419-024-07117-3. Cell Death Dis. 2024. PMID: 39500888 Free PMC article.
-
Chromatin Dynamics in Digestive System Cancer: Commander and Regulator.Front Oncol. 2022 Jul 29;12:935877. doi: 10.3389/fonc.2022.935877. eCollection 2022. Front Oncol. 2022. PMID: 35965507 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

