Plant E3 Ligases and Their Role in Abiotic Stress Response
- PMID: 35269512
- PMCID: PMC8909703
- DOI: 10.3390/cells11050890
Plant E3 Ligases and Their Role in Abiotic Stress Response
Abstract
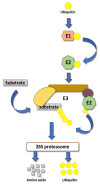
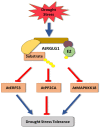
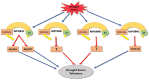
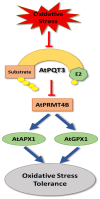
Plants, as sessile organisms, have limited means to cope with environmental changes. Consequently, they have developed complex regulatory systems to ameliorate abiotic stresses im-posed by environmental changes. One such system is the ubiquitin proteasome pathway, which utilizes E3 ligases to target proteins for proteolytic degradation via the 26S proteasome. Plants ex-press a plethora of E3 ligases that are categorized into four major groups depending on their structure. They are involved in many biological and developmental processes in plants, such as DNA repair, photomorphogenesis, phytohormones signaling, and biotic stress. Moreover, many E3 ligase targets are proteins involved in abiotic stress responses, such as salt, drought, heat, and cold. In this review, we will provide a comprehensive overview of E3 ligases and their substrates that have been connected with abiotic stress in order to illustrate the diversity and complexity of how this pathway enables plant survival under stress conditions.
Keywords: E3 ligase; abiotic stress; drought stress; heavy metal stress; oxidative stress; salt stress; temperature stress.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


Similar articles
-
A review on ubiquitin ligases: Orchestrators of plant resilience in adversity.Plant Sci. 2024 Oct;347:112180. doi: 10.1016/j.plantsci.2024.112180. Epub 2024 Jul 2. Plant Sci. 2024. PMID: 38964613 Review.
-
Small ubiquitin-like modifiers E3 ligases in plant stress.Funct Plant Biol. 2024 Apr;51:FP24032. doi: 10.1071/FP24032. Funct Plant Biol. 2024. PMID: 38669463 Review.
-
Transcriptome Profiling and Functional Validation of RING-Type E3 Ligases in Halophyte Sesuvium verrucosum under Salinity Stress.Int J Mol Sci. 2022 Mar 4;23(5):2821. doi: 10.3390/ijms23052821. Int J Mol Sci. 2022. PMID: 35269961 Free PMC article.
-
Oryza sativa drought-, heat-, and salt-induced RING finger protein 1 (OsDHSRP1) negatively regulates abiotic stress-responsive gene expression.Plant Mol Biol. 2020 Jun;103(3):235-252. doi: 10.1007/s11103-020-00989-x. Epub 2020 Mar 21. Plant Mol Biol. 2020. PMID: 32206999
-
Roles of E3 Ubiquitin Ligases in Plant Responses to Abiotic Stresses.Int J Mol Sci. 2022 Feb 19;23(4):2308. doi: 10.3390/ijms23042308. Int J Mol Sci. 2022. PMID: 35216424 Free PMC article. Review.
Cited by
-
An insight into the roles of ubiquitin-specific proteases in plants: development and growth, morphogenesis, and stress response.Front Plant Sci. 2024 Jun 27;15:1396634. doi: 10.3389/fpls.2024.1396634. eCollection 2024. Front Plant Sci. 2024. PMID: 38993940 Free PMC article. Review.
-
The F-box protein ZmFBL41 negatively regulates disease resistance to Rhizoctonia solani by degrading the abscisic acid synthase ZmNCED6 in maize.Plant Cell Rep. 2024 Feb 1;43(2):48. doi: 10.1007/s00299-023-03132-4. Plant Cell Rep. 2024. PMID: 38300347
-
QTL mapping for yield contributing traits in mungbean (Vigna radiata L.) using a RIL population.Sci Rep. 2025 Jul 1;15(1):20795. doi: 10.1038/s41598-025-99687-1. Sci Rep. 2025. PMID: 40596720 Free PMC article.
-
Genome-Wide Identification and Functional Analysis of the Genes of the ATL Family in Maize during High-Temperature Stress in Maize.Genes (Basel). 2024 Aug 22;15(8):1106. doi: 10.3390/genes15081106. Genes (Basel). 2024. PMID: 39202465 Free PMC article.
-
Genome-wide identification and expression profiling reveal the regulatory role of U-box E3 ubiquitin ligase genes in strawberry fruit ripening and abiotic stresses resistance.Front Plant Sci. 2023 Mar 23;14:1171056. doi: 10.3389/fpls.2023.1171056. eCollection 2023. Front Plant Sci. 2023. PMID: 37035055 Free PMC article.
References
-
- Kajla M., Yadav V.K., Khokhar J., Singh S., Chhokar R., Meena R.P., Sharma R. Increase in wheat production through management of abiotic stresses: A review. J. Appl. Nat. Sci. 2015;7:1070–1080. doi: 10.31018/jans.v7i2.733. - DOI
-
- Iizumi T., Shen Z., Furuya J., Koizumi T., Furuhashi G., Kim W., Nishimori M. Climate change adaptation cost and residual damage to global crop production. Clim. Res. 2020;80:203–218. doi: 10.3354/cr01605. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

