Mechanism Underlying the Bypass of Apurinic/Pyrimidinic Site Analogs by Sulfolobus acidocaldarius DNA Polymerase IV
- PMID: 35269871
- PMCID: PMC8910976
- DOI: 10.3390/ijms23052729
Mechanism Underlying the Bypass of Apurinic/Pyrimidinic Site Analogs by Sulfolobus acidocaldarius DNA Polymerase IV
Abstract
The spontaneous depurination of genomic DNA occurs frequently and generates apurinic/pyrimidinic (AP) site damage that is mutagenic or lethal to cells. Error-prone DNA polymerases are specifically responsible for the translesion synthesis (TLS) of specific DNA damage, such as AP site damage, generally with relatively low fidelity. The Y-family DNA polymerases are the main error-prone DNA polymerases, and they employ three mechanisms to perform TLS, including template-skipping, dNTP-stabilized misalignment, and misincorporation-misalignment. The bypass mechanism of the dinB homolog (Dbh), an archaeal Y-family DNA polymerase from Sulfolobus acidocaldarius, is unclear and needs to be confirmed. In this study, we show that the Dbh primarily uses template skipping accompanied by dNTP-stabilized misalignment to bypass AP site analogs, and the incorporation of the first nucleotide across the AP site is the most difficult. Furthermore, based on the reported crystal structures, we confirmed that three conserved residues (Y249, R333, and I295) in the little finger (LF) domain and residue K78 in the palm subdomain of the catalytic core domain are very important for TLS. These results deepen our understanding of how archaeal Y-family DNA polymerases deal with intracellular AP site damage and provide a biochemical basis for elucidating the intracellular function of these polymerases.
Keywords: AP site analogs; Dbh; Sulfolobus acidocaldarius; little finger domain; translesion synthesis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
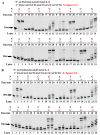
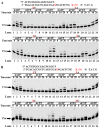



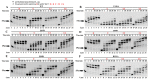
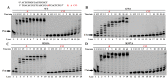
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

