mt tRFs, New Players in MELAS Disease
- PMID: 35273517
- PMCID: PMC8902416
- DOI: 10.3389/fphys.2022.800171
mt tRFs, New Players in MELAS Disease
Abstract
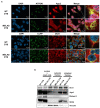
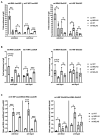
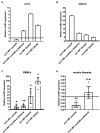
MELAS (mitochondrial encephalomyopathy, lactic acidosis, and stroke-like episodes) is an OXPHOS disease mostly caused by the m.3243A>G mutation in the mitochondrial tRNALeu(UUR) gene. Recently, we have shown that the mutation significantly changes the expression pattern of several mitochondrial tRNA-derived small RNAs (mt tsRNAs or mt tRFs) in a cybrid model of MELAS and in fibroblasts from MELAS patients versus control cells. Among them are those derived from mt tRNA LeuUUR containing or not the m.3243A>G mutation (mt 5'-tRF LeuUUR-m.3243A>G and mt 5'-tRF LeuUUR), whose expression levels are, respectively, increased and decreased in both MELAS cybrids and fibroblasts. Here, we asked whether mt 5'-tRF LeuUUR and mt 5'-tRF LeuUUR-m.3243A>G are biologically relevant and whether these mt tRFs are detected in diverse patient samples. Treatment with a mimic oligonucleotide of mt tRNA LeuUUR fragment (mt 5'-tRF LeuUUR) showed a therapeutic potential since it partially restored mitochondrial respiration in MELAS cybrids. Moreover, these mt tRFs could be detected in biofluids like urine and blood. We also investigated the participation of miRNA pathway components Dicer and Ago2 in the mt tRFs biogenesis process. We found that Dicer and Ago2 localize in the mitochondria of MELAS cybrids and that immunoprecipitation of these proteins in cytoplasm and mitochondria fractions revealed an increased mt tRF/mt tRNA ratio in MELAS condition compared to WT. These preliminary results suggest an involvement of Dicer and Ago2 in the mechanism of mt tRF biogenesis and action.
Keywords: mitochondrial dysfunction; retrograde signaling; sncRNAs; tRF and tiRNA; tRNA fragment.
Copyright © 2022 Meseguer and Rubio.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Bose M., Chatterjee S., Chakrabarty Y., Barman B., Bhattacharyya S. N. (2020). Retrograde trafficking of Argonaute 2 acts as a rate-limiting step for de novo miRNP formation on endoplasmic reticulum-attached polysomes in mammalian cells. Life Sci. Alliance. 3:e201800161. doi: 10.26508/lsa.201800161, PMID: - DOI - PMC - PubMed
-
- Burroughs A. M., Ando Y., de Hoon M. J., Tomaru Y., Suzuki H., Hayashizaki Y., et al. . (2011). Deep-sequencing of human Argonaute-associated small RNAs provides insight into miRNA sorting and reveals Argonaute association with RNA fragments of diverse origin. RNA Biol. 8, 158–177. doi: 10.4161/rna.8.1.14300, PMID: - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

