A Quantitative Approach to Unravel the Role of Host Genetics in IgG-FcγR Complex Formation After Vaccination
- PMID: 35273603
- PMCID: PMC8902241
- DOI: 10.3389/fimmu.2022.820148
A Quantitative Approach to Unravel the Role of Host Genetics in IgG-FcγR Complex Formation After Vaccination
Abstract
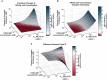
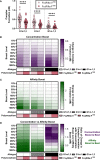
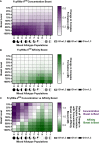
Fc-mediated immune functions have been correlated with protection in the RV144 HIV vaccine trial and are important for immunity to a range of pathogens. IgG antibodies (Abs) that form complexes with Fc receptors (FcRs) on innate immune cells can activate Fc-mediated immune functions. Genetic variation in both IgGs and FcRs have the capacity to alter IgG-FcR complex formation via changes in binding affinity and concentration. A growing challenge lies in unraveling the importance of multiple variations, especially in the context of vaccine trials that are conducted in homogenous genetic populations. Here we use an ordinary differential equation model to quantitatively assess how IgG1 allotypes and FcγR polymorphisms influence IgG-FcγRIIIa complex formation in vaccine-relevant settings. Using data from the RV144 HIV vaccine trial, we map the landscape of IgG-FcγRIIIa complex formation predicted post-vaccination for three different IgG1 allotypes and two different FcγRIIIa polymorphisms. Overall, the model illustrates how specific vaccine interventions could be applied to maximize IgG-FcγRIIIa complex formation in different genetic backgrounds. Individuals with the G1m1,17 and G1m1,3 allotypes were predicted to be more responsive to vaccine adjuvant strategies that increase antibody FcγRIIIa affinity (e.g. glycosylation modifications), compared to the G1m-1,3 allotype which was predicted to be more responsive to vaccine boosting regimens that increase IgG1 antibody titers (concentration). Finally, simulations in mixed-allotype populations suggest that the benefit of boosting IgG1 concentration versus IgG1 affinity may be dependent upon the presence of the G1m-1,3 allotype. Overall this work provides a quantitative tool for rationally improving Fc-mediated functions after vaccination that may be important for assessing vaccine trial results in the context of under-represented genetic populations.
Keywords: ADCC; Fc receptor; Fc receptor polymorphism; HIV; IgG1 allotype; RV144; systems serology; vaccine boosting.
Copyright © 2022 Lemke, Theisen, Bozich, McLean, Lee, Lopez, Rerks-Ngarm, Pitisuttithum, Nitayaphan, Kratochvil, Wines, Hogarth, Kent, Chung and Arnold.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

