Gene Regulatory Networks of Epidermal and Neural Fate Choice in a Chordate
- PMID: 35276009
- PMCID: PMC9004418
- DOI: 10.1093/molbev/msac055
Gene Regulatory Networks of Epidermal and Neural Fate Choice in a Chordate
Abstract
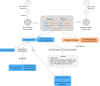
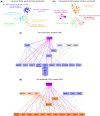
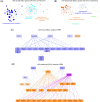
Neurons are a highly specialized cell type only found in metazoans. They can be scattered throughout the body or grouped together, forming ganglia or nerve cords. During embryogenesis, centralized nervous systems develop from the ectoderm, which also forms the epidermis. How pluripotent ectodermal cells are directed toward neural or epidermal fates, and to which extent this process is shared among different animal lineages, are still open questions. Here, by using micromere explants, we were able to define in silico the putative gene regulatory networks (GRNs) underlying the first steps of the epidermis and the central nervous system formation in the cephalochordate amphioxus. We propose that although the signal triggering neural induction in amphioxus (i.e., Nodal) is different from vertebrates, the main transcription factors implicated in this process are conserved. Moreover, our data reveal that transcription factors of the neural program seem to not only activate neural genes but also to potentially have direct inputs into the epidermal GRN, suggesting that the Nodal signal might also contribute to neural fate commitment by repressing the epidermal program. Our functional data on whole embryos support this result and highlight the complex interactions among the transcription factors activated by the signaling pathways that drive ectodermal cell fate choice in chordates.
Keywords: EvoDevo; amphioxus; cell fate; gene regulatory network; neural induction.
© The Author(s) 2022. Published by Oxford University Press on behalf of Society for Molecular Biology and Evolution.
Figures



Similar articles
-
Essential role of Bmp signaling and its positive feedback loop in the early cell fate evolution of chordates.Dev Biol. 2013 Oct 15;382(2):538-54. doi: 10.1016/j.ydbio.2013.07.021. Epub 2013 Aug 7. Dev Biol. 2013. PMID: 23933491
-
The choice between epidermal and neural fate: a matter of calcium.Int J Dev Biol. 2004;48(2-3):75-84. doi: 10.1387/ijdb.15272372. Int J Dev Biol. 2004. PMID: 15272372 Review.
-
Evolutionary origin of the chordate nervous system revealed by amphioxus developmental trajectories.Nat Ecol Evol. 2024 Sep;8(9):1693-1710. doi: 10.1038/s41559-024-02469-7. Epub 2024 Jul 18. Nat Ecol Evol. 2024. PMID: 39025981
-
Highly distinct genetic programs for peripheral nervous system formation in chordates.BMC Biol. 2022 Jun 27;20(1):152. doi: 10.1186/s12915-022-01355-7. BMC Biol. 2022. PMID: 35761237 Free PMC article.
-
Enhancer evolution in chordates: Lessons from functional analyses of cephalochordate cis-regulatory modules.Dev Growth Differ. 2020 Jun;62(5):279-300. doi: 10.1111/dgd.12684. Epub 2020 Jun 16. Dev Growth Differ. 2020. PMID: 32479656 Review.
Cited by
-
Chromatin Landscape Governing Murine Epidermal Differentiation.J Invest Dermatol. 2023 Jul;143(7):1220-1232.e9. doi: 10.1016/j.jid.2022.12.020. Epub 2023 Jan 26. J Invest Dermatol. 2023. PMID: 36708949 Free PMC article.
-
An amphioxus neurula stage cell atlas supports a complex scenario for the emergence of vertebrate head mesoderm.Nat Commun. 2024 May 29;15(1):4550. doi: 10.1038/s41467-024-48774-4. Nat Commun. 2024. PMID: 38811547 Free PMC article.
-
Retinoic Acid and POU Genes in Developing Amphioxus: A Focus on Neural Development.Cells. 2023 Feb 14;12(4):614. doi: 10.3390/cells12040614. Cells. 2023. PMID: 36831281 Free PMC article.
References
-
- Albuixech-Crespo B, Lopez-Blanch L, Burguera D, Maeso I, Sanchez-Arrones L, Moreno-Bravo JA, Somorjai I, Pascual-Anaya J, Puelles E, Bovolenta P, et al. . 2017. Molecular regionalization of the developing amphioxus neural tube challenges major partitions of the vertebrate brain. PLoS Biol. 15:e2001573. - PMC - PubMed
-
- Aldea D, Subirana L, Keime C, Meister L, Maeso I, Marcellini S, Gomez-Skarmeta JL, Bertrand S, Escriva H. 2019. Genetic regulation of amphioxus somitogenesis informs the evolution of the vertebrate head mesoderm. Nat Ecol Evol. 3:1233–1240. - PubMed
-
- Arendt D, Nubler-Jung K. 1999. Comparison of early nerve cord development in insects and vertebrates. Development 126:2309–2325. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

