Computational models for studying physical instabilities in high concentration biotherapeutic formulations
- PMID: 35282775
- PMCID: PMC8928847
- DOI: 10.1080/19420862.2022.2044744
Computational models for studying physical instabilities in high concentration biotherapeutic formulations
Abstract
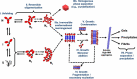
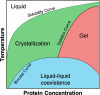
Computational prediction of the behavior of concentrated protein solutions is particularly advantageous in early development stages of biotherapeutics when material availability is limited and a large set of formulation conditions needs to be explored. This review provides an overview of the different computational paradigms that have been successfully used in modeling undesirable physical behaviors of protein solutions with a particular emphasis on high-concentration drug formulations. This includes models ranging from all-atom simulations, coarse-grained representations to macro-scale mathematical descriptions used to study physical instability phenomena of protein solutions such as aggregation, elevated viscosity, and phase separation. These models are compared and summarized in the context of the physical processes and their underlying assumptions and limitations. A detailed analysis is also given for identifying protein interaction processes that are explicitly or implicitly considered in the different modeling approaches and particularly their relations to various formulation parameters. Lastly, many of the shortcomings of existing computational models are discussed, providing perspectives and possible directions toward an efficient computational framework for designing effective protein formulations.
Keywords: Biotherapeutics; aggregation; drug formulation; high concentration; molecular modeling; phase separation; physical instabilities; viscosity.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures


References
-
- Warne NW, Mahler H-C. Challenges in protein product development. Cham, Switzerland: Springer International Publishing; 2018.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
