Target-oriented prioritization: targeted selection strategy by integrating organismal and molecular traits through predictive analytics in breeding
- PMID: 35292095
- PMCID: PMC8922918
- DOI: 10.1186/s13059-022-02650-w
Target-oriented prioritization: targeted selection strategy by integrating organismal and molecular traits through predictive analytics in breeding
Abstract
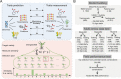
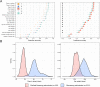
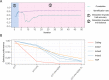
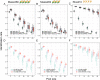
Genomic prediction in crop breeding is hindered by modeling on limited phenotypic traits. We propose an integrative multi-trait breeding strategy via machine learning algorithm, target-oriented prioritization (TOP). Using a large hybrid maize population, we demonstrate that the accuracy for identifying a candidate that is phenotypically closest to an ideotype, or target variety, achieves up to 91%. The strength of TOP is enhanced when omics level traits are included. We show that TOP enables selection of inbreds or hybrids that outperform existing commercial varieties. It improves multiple traits and accurately identifies improved candidates for new varieties, which will greatly influence breeding.
Keywords: Crop breeding; Genomic prediction; Machine learning; Multiple traits; Omics.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Genomic Prediction of Complex Traits in an Allogamous Annual Crop: The Case of Maize Single-Cross Hybrids.Methods Mol Biol. 2022;2467:543-567. doi: 10.1007/978-1-0716-2205-6_20. Methods Mol Biol. 2022. PMID: 35451790
-
Genomic prediction applied to multiple traits and environments in second season maize hybrids.Heredity (Edinb). 2020 Aug;125(1-2):60-72. doi: 10.1038/s41437-020-0321-0. Epub 2020 May 29. Heredity (Edinb). 2020. PMID: 32472060 Free PMC article.
-
Beyond Genomic Prediction: Combining Different Types of omics Data Can Improve Prediction of Hybrid Performance in Maize.Genetics. 2018 Apr;208(4):1373-1385. doi: 10.1534/genetics.117.300374. Epub 2018 Jan 23. Genetics. 2018. PMID: 29363551 Free PMC article.
-
Machine learning bridges omics sciences and plant breeding.Trends Plant Sci. 2023 Feb;28(2):199-210. doi: 10.1016/j.tplants.2022.08.018. Epub 2022 Sep 21. Trends Plant Sci. 2023. PMID: 36153276 Review.
-
Integrating omics databases for enhanced crop breeding.J Integr Bioinform. 2023 Jul 25;20(4):20230012. doi: 10.1515/jib-2023-0012. eCollection 2023 Dec 1. J Integr Bioinform. 2023. PMID: 37486120 Free PMC article. Review.
Cited by
-
CropGS-Hub: a comprehensive database of genotype and phenotype resources for genomic prediction in major crops.Nucleic Acids Res. 2024 Jan 5;52(D1):D1519-D1529. doi: 10.1093/nar/gkad1062. Nucleic Acids Res. 2024. PMID: 38000385 Free PMC article.
-
Integration of multi-omics technologies for crop improvement: Status and prospects.Front Bioinform. 2022 Oct 19;2:1027457. doi: 10.3389/fbinf.2022.1027457. eCollection 2022. Front Bioinform. 2022. PMID: 36438626 Free PMC article.
-
Metabolic marker-assisted genomic prediction improves hybrid breeding.Plant Commun. 2025 Mar 10;6(3):101199. doi: 10.1016/j.xplc.2024.101199. Epub 2024 Nov 29. Plant Commun. 2025. PMID: 39614617 Free PMC article.
-
Adaptive and maladaptive introgression in grapevine domestication.Proc Natl Acad Sci U S A. 2023 Jun 13;120(24):e2222041120. doi: 10.1073/pnas.2222041120. Epub 2023 Jun 5. Proc Natl Acad Sci U S A. 2023. PMID: 37276420 Free PMC article.
-
Machine Learning for AI Breeding in Plants.Genomics Proteomics Bioinformatics. 2024 Sep 13;22(4):qzae051. doi: 10.1093/gpbjnl/qzae051. Genomics Proteomics Bioinformatics. 2024. PMID: 38954837 Free PMC article. No abstract available.
References
-
- Steinwand MA, Ronald PC. Crop biotechnology and the future of food. Nat Food. 2020;1(5):273–283.
-
- Hickey JM, Chiurugwi T, Mackay I, Powell W, Eggen A, Kilian A, Jones C, Canales C, Grattapaglia D, Bassi F. Genomic prediction unifies animal and plant breeding programs to form platforms for biological discovery. Nat Genet. 2017;49(9):1297. - PubMed
-
- Hickey LT, Hafeez AN, Robinson H, Jackson SA, Leal-Bertioli SC, Tester M, Gao C, Godwin ID, Hayes BJ, Wulff BB. Breeding crops to feed 10 billion. Nat Biotechnol. 2019;37(7):744–754. - PubMed
-
- Borlaug NE. Contributions of conventional plant breeding to food production. Science. 1983;219(4585):689–693. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

