Kinetic interplay between droplet maturation and coalescence modulates shape of aged protein condensates
- PMID: 35293386
- PMCID: PMC8924231
- DOI: 10.1038/s41598-022-08130-2
Kinetic interplay between droplet maturation and coalescence modulates shape of aged protein condensates
Abstract
Biomolecular condensates formed by the process of liquid-liquid phase separation (LLPS) play diverse roles inside cells, from spatiotemporal compartmentalisation to speeding up chemical reactions. Upon maturation, the liquid-like properties of condensates, which underpin their functions, are gradually lost, eventually giving rise to solid-like states with potential pathological implications. Enhancement of inter-protein interactions is one of the main mechanisms suggested to trigger the formation of solid-like condensates. To gain a molecular-level understanding of how the accumulation of stronger interactions among proteins inside condensates affect the kinetic and thermodynamic properties of biomolecular condensates, and their shapes over time, we develop a tailored coarse-grained model of proteins that transition from establishing weak to stronger inter-protein interactions inside condensates. Our simulations reveal that the fast accumulation of strongly binding proteins during the nucleation and growth stages of condensate formation results in aspherical solid-like condensates. In contrast, when strong inter-protein interactions appear only after the equilibrium condensate has been formed, or when they accumulate slowly over time with respect to the time needed for droplets to fuse and grow, spherical solid-like droplets emerge. By conducting atomistic potential-of-mean-force simulations of NUP-98 peptides-prone to forming inter-protein [Formula: see text]-sheets-we observe that formation of inter-peptide [Formula: see text]-sheets increases the strength of the interactions consistently with the loss of liquid-like condensate properties we observe at the coarse-grained level. Overall, our work aids in elucidating fundamental molecular, kinetic, and thermodynamic mechanisms linking the rate of change in protein interaction strength to condensate shape and maturation during ageing.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



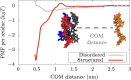
References
-
- Sear RP. The cytoplasm of living cells: A functional mixture of thousands of components. J. Phys. Condens. Matter. 2005;17:S3587–S3595.
-
- Shin Y, Brangwynne CP. Liquid phase condensation in cell physiology and disease. Science. 2017;357:eaaf4382. - PubMed
-
- Alberts B. Molecular Biology of the Cell. 6. New York, NY: Garland Science, Taylor and Francis Group; 2015.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

