Photoaffinity Capture Compounds to Profile the Magic Spot Nucleotide Interactomes
- PMID: 35294098
- PMCID: PMC9310846
- DOI: 10.1002/anie.202201731
Photoaffinity Capture Compounds to Profile the Magic Spot Nucleotide Interactomes
Abstract
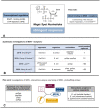
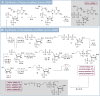
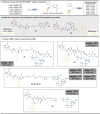
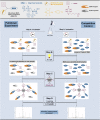
Magic Spot Nucleotides (MSN) regulate the stringent response, a highly conserved bacterial stress adaptation mechanism, enabling survival under adverse external challenges. In times of antibiotic crisis, a detailed understanding of stringent response is essential, as potentially new targets for pharmacological intervention could be identified. In this study, we delineate the MSN interactome in Escherichia coli and Salmonella typhimurium applying a family of trifunctional photoaffinity capture compounds. We introduce MSN probes covering a diverse phosphorylation pattern, such as pppGpp, ppGpp, and pGpp. Our chemical proteomics approach provides datasets of putative MSN receptors both from cytosolic and membrane fractions that unveil new MSN targets. We find that the activity of the non-Nudix hydrolase ApaH is potently inhibited by pppGpp, which itself is converted to pGpp by ApaH. The capture compounds described herein will be useful to identify MSN interactomes across bacterial species.
Keywords: Alarmones; Magic Spot Nucleotides; Photoaffinity Proteomics; Stringent Response; ppGpp.
© 2022 The Authors. Angewandte Chemie International Edition published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

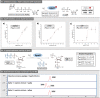
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

