Beacon v2 and Beacon networks: A "lingua franca" for federated data discovery in biomedical genomics, and beyond
- PMID: 35297548
- PMCID: PMC9322265
- DOI: 10.1002/humu.24369
Beacon v2 and Beacon networks: A "lingua franca" for federated data discovery in biomedical genomics, and beyond
Abstract
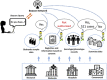
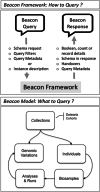
Beacon is a basic data discovery protocol issued by the Global Alliance for Genomics and Health (GA4GH). The main goal addressed by version 1 of the Beacon protocol was to test the feasibility of broadly sharing human genomic data, through providing simple "yes" or "no" responses to queries about the presence of a given variant in datasets hosted by Beacon providers. The popularity of this concept has fostered the design of a version 2, that better serves real-world requirements and addresses the needs of clinical genomics research and healthcare, as assessed by several contributing projects and organizations. Particularly, rare disease genetics and cancer research will benefit from new case level and genomic variant level requests and the enabling of richer phenotype and clinical queries as well as support for fuzzy searches. Beacon is designed as a "lingua franca" to bridge data collections hosted in software solutions with different and rich interfaces. Beacon version 2 works alongside popular standards like Phenopackets, OMOP, or FHIR, allowing implementing consortia to return matches in beacon responses and provide a handover to their preferred data exchange format. The protocol is being explored by other research domains and is being tested in several international projects.
Keywords: Beacon; GA4GH; REST API; clinical genomics; data discovery; data sharing.
© 2022 The Authors. Human Mutation published by Wiley Periodicals LLC.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
References
-
- Ayaz, M. , Pasha, M. F. , Alzahrani, M. Y. , Budiarto, R. , & Stiawan, D. (2021). The fast health interoperability resources (FHIR) standard: Systematic literature review of implementations, applications, challenges and opportunities. JMIR Medical Informatics, 9(7), e21929. 10.2196/21929 - DOI - PMC - PubMed
-
- Bosca, D. , Moner, D. , Maldonado, J. A. , & Robles, M. (2015). Combining archetypes with fast health interoperability resources in future‐proof health information systems. Studies in Health Technology and Informatics, 210, 180–184. - PubMed
-
- Cezard, T. , Cunningham, F. , Hunt, S. E. , Koylass, B. , Kumar, N. , Saunders, G. , Shen, A. , Silva, A. F. , Tsukanov, K. , Venkataraman, S. , Flicek, P. , Parkinson, H. , & Keane, T. M. (2022). The European Variation Archive: A FAIR resource of genomic variation for all species. Nucleic Acids Research, 50, gkab960. 10.1093/nar/gkab960 - DOI - PMC - PubMed
-
- Cline, M. S. , Liao, R. G. , Parsons, M. T. , Paten, B. , Alquaddoomi, F. , Antoniou, A. , Baxter, S. , Brody, L. , Cook‐Deegan, R. , Coffin, A. , Couch, F. J. , Craft, B. , Currie, R. , Dlott, C. C. , Dolman, L. , den Dunnen, J. T. , Dyke, S. , Domchek, S. M. , Easton, D. , … Spurdle, A. B. (2018). BRCA Challenge: BRCA Exchange as a global resource for variants in BRCA1 and BRCA2. PLoS Genetics, 14, e1007752. 10.1371/journal.pgen.1007752 - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

