Role of chromatin modulator Dpy30 in osteoclast differentiation and function
- PMID: 35307321
- PMCID: PMC9063347
- DOI: 10.1016/j.bone.2022.116379
Role of chromatin modulator Dpy30 in osteoclast differentiation and function
Abstract
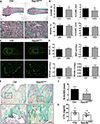
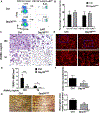
Osteoclasts are the principal bone resorption cells crucial for homeostatic bone remodeling and pathological bone destruction. Increasing data demonstrate a vital role of histone methylation in osteoclastogenesis. As an integral core subunit of H3K4 methyltransferases, Dpy30 is notal as a key chromatin regulator for cell growth and differentiation and stem cell fate determination, particularly in the hematopoietic system. However, its role in osteoclastogenesis is currently unknown. Herein, we generated Dpy30F/F; LysM-Cre+/+ mice, which deletes Dpy30 in myeloid cells, to characterize its involvement in osteoclast differentiation and function. Dpy30F/F; LysM-Cre+/+ mice showed increased bone mass, evident by impaired osteoclastogenesis and defective osteoclast activity, but no alteration of osteoblast numbers and bone formation. Additionally, our ex vivo analysis showed that the loss of Dpy30 significantly impedes osteoclast differentiation and suppresses osteoclast-related gene expression. Moreover, Dpy30 deficiency significantly decreased the enrichment of H3K4me3 on the promoter region of NFATc1. Thus, we revealed a novel role for Dpy30 in osteoclastogenesis through epigenetic mechanisms, and that it could potentially be a therapeutic target for bone destruction diseases.
Keywords: Bone resorption; Dpy30; H3K4 methylation; Osteoclast.
Copyright © 2022 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of competing interest
The authors declare no conflicts of interest.
Figures
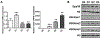

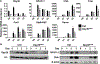

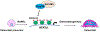
References
-
- Boyle WJ, Simonet WS, and Lacey DL (2003) Osteoclast differentiation and activation. Nature 423, 337–342 - PubMed
-
- Novack DV, and Teitelbaum SL (2008) The osteoclast: friend or foe? Annu Rev Pathol 3, 457–484 - PubMed
-
- Teitelbaum SL, and Ross FP (2003) Genetic regulation of osteoclast development and function. Nat Rev Genet 4, 638–649 - PubMed
-
- Takayanagi H, Kim S, Koga T, Nishina H, Isshiki M, Yoshida H, Saiura A, Isobe M, Yokochi T, Inoue J, Wagner EF, Mak TW, Kodama T, and Taniguchi T (2002) Induction and activation of the transcription factor NFATc1 (NFAT2) integrate RANKL signaling in terminal differentiation of osteoclasts. Dev Cell 3, 889–901 - PubMed
-
- Takeshita S, Kaji K, and Kudo A (2000) Identification and characterization of the new osteoclast progenitor with macrophage phenotypes being able to differentiate into mature osteoclasts. J Bone Miner Res 15, 1477–1488 - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

