Modeling Human Primary Microcephaly With hiPSC-Derived Brain Organoids Carrying CPAP-E1235V Disease-Associated Mutant Protein
- PMID: 35309908
- PMCID: PMC8924525
- DOI: 10.3389/fcell.2022.830432
Modeling Human Primary Microcephaly With hiPSC-Derived Brain Organoids Carrying CPAP-E1235V Disease-Associated Mutant Protein
Abstract
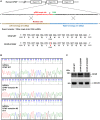
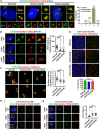
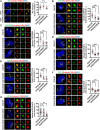
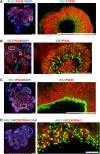
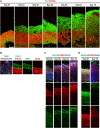
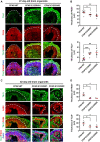
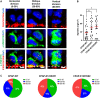
The centrosome is composed of a pair of centrioles and serves as the major microtubule-organizing center (MTOC) in cells. Centrosome dysfunction has been linked to autosomal recessive primary microcephaly (MCPH), which is a rare human neurodevelopmental disorder characterized by small brain size with intellectual disability. Recently, several mouse models carrying mutated genes encoding centrosomal proteins have been generated to address the genotype-phenotype relationships in MCPH. However, several human-specific features were not observed in the mouse models during brain development. Herein, we generated isogenic hiPSCs carrying the gene encoding centrosomal CPAP-E1235V mutant protein using the CRISPR-Cas9 genome editing system, and examined the phenotypic features of wild-type and mutant hiPSCs and their derived brain organoids. Our results showed that the CPAP-E1235V mutant perturbed the recruitment of several centriolar proteins involved in centriole elongation, including CEP120, CEP295, CENTROBIN, POC5, and POC1B, onto nascent centrioles, resulting in the production of short centrioles but long cilia. Importantly, our wild-type hiPSC-derived brain organoid recapitulated many cellular events seen in the developing human brain, including neuronal differentiation and cortical spatial lamination. Interestingly, hiPSC-CPAP-E1235V-derived brain organoids induced p53-dependent neuronal cell death, resulting in the production of smaller brain organoids that mimic the microcephaly phenotype. Furthermore, we observed that the CPAP-E1235V mutation altered the spindle orientation of neuronal progenitor cells and induced premature neuronal differentiation. In summary, we have shown that the hiPSC-derived brain organoid coupled with CRISPR/Cas9 gene editing technology can recapitulate the centrosome/centriole-associated MCPH pathological features. Possible mechanisms for MCPH with centriole/centrosome dysfunction are discussed.
Keywords: CENPJ; CPAP; centriole duplication; cerebral organoid model; ciliopathy; human microcephaly; microcephaly gene mutations.
Copyright © 2022 An, Kuo and Tang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

