Identification of N6-Methyladenosine-Related lncRNAs as a Prognostic Signature in Glioma
- PMID: 35311131
- PMCID: PMC8927984
- DOI: 10.3389/fonc.2022.789283
Identification of N6-Methyladenosine-Related lncRNAs as a Prognostic Signature in Glioma
Abstract
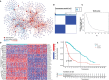
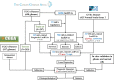
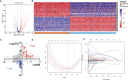
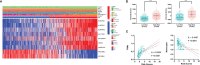
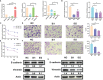
N6-methyladenosine (m6A) modification is the most abundant modification in long noncoding RNAs (lncRNAs). Current studies have shown that the abnormal expression of m6A-related genes is closely associated with the tumorigenesis and progression of glioma. However, the role of m6A-related lncRNAs in glioma development is still unclear. Herein, we screened 566 m6A-related lncRNAs in glioma from The Cancer Genome Atlas (TCGA) database. The expression pattern of these lncRNAs could cluster samples into two groups, in which various classical tumor-related functions and the tumor immune microenvironment were significantly different. Subsequently, a nine-factor m6A-related lncRNA prognostic signature (MLPS) was constructed by using a LASSO regression analysis in the training set and was validated in the test set and independent datasets. The AUC values of the MLPS were 0.881, 0.918 and 0.887 for 1-, 3- and 5-year survival in the training set, respectively, and 0.856, 0.916 and 0.909 for 1-, 3-, and 5-year survival in the test set, respectively. Stratification analyses of the MLPS illustrated its prognostic performance in gliomas with different characteristics. Correlation analyses showed that the infiltrations of monocytes and tumor-associated macrophages (TAMs) were significantly relevant to the risk score in the MLPS. Moreover, we detected the expression of four MLPS factors with defined sequences in glioma and normal cells by using RT-PCR. Afterwards, we investigated the functions of LNCTAM34A (one of the MLPS factors) in glioma cells, which have rarely been reported. Via in vitro experiments, LNCTAM34A was demonstrated to promote the proliferation, migration and epithelial-mesenchymal transition (EMT) of glioma cells. Overall, our study revealed the critical role of m6A-related lncRNAs in glioma and elucidated that LNCTAM34A could promote glioma proliferation, migration and EMT.
Keywords: N6-methyladenosine; The Cancer Genome Atlas; glioma; long non-coding RNA; prognostic signature.
Copyright © 2022 Chen, Guo, Li, Xu, Wang, Ning, Ma, Qu, Zhang and Zhang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
The prognostic value and immune landscaps of m6A/m5C-related lncRNAs signature in the low grade glioma.BMC Bioinformatics. 2023 Jul 4;24(1):274. doi: 10.1186/s12859-023-05386-x. BMC Bioinformatics. 2023. PMID: 37403043 Free PMC article.
-
N6-methyladenosine-related lncRNAs identified as potential biomarkers for predicting the overall survival of Asian gastric cancer patients.BMC Cancer. 2022 Jul 1;22(1):721. doi: 10.1186/s12885-022-09801-z. BMC Cancer. 2022. PMID: 35778697 Free PMC article.
-
Comprehensive analyses indicated the association between m6A related long non-coding RNAs and various pathways in glioma.Cancer Med. 2023 Jan;12(1):760-788. doi: 10.1002/cam4.4913. Epub 2022 Jun 6. Cancer Med. 2023. PMID: 35668574 Free PMC article.
-
The functions of N6-methyladenosine modification in lncRNAs.Genes Dis. 2020 Mar 19;7(4):598-605. doi: 10.1016/j.gendis.2020.03.005. eCollection 2020 Dec. Genes Dis. 2020. PMID: 33335959 Free PMC article. Review.
-
Research Progress on Long Noncoding RNAs and N6-Methyladenosine in Hepatocellular Carcinoma.Front Oncol. 2022 Jul 22;12:907399. doi: 10.3389/fonc.2022.907399. eCollection 2022. Front Oncol. 2022. PMID: 35936671 Free PMC article. Review.
Cited by
-
MBNL1‑AS1 attenuates tumor cell proliferation by regulating the miR‑29c‑3p/BVES signal in colorectal cancer.Oncol Rep. 2023 Oct;50(4):191. doi: 10.3892/or.2023.8628. Epub 2023 Sep 15. Oncol Rep. 2023. PMID: 37711058 Free PMC article.
-
An m6A/m5C/m1A/m7G-Related Long Non-coding RNA Signature to Predict Prognosis and Immune Features of Glioma.Front Genet. 2022 May 26;13:903117. doi: 10.3389/fgene.2022.903117. eCollection 2022. Front Genet. 2022. PMID: 35692827 Free PMC article.
-
Gene signature of m6A RNA regulators in diagnosis, prognosis, treatment, and immune microenvironment for cervical cancer.Sci Rep. 2022 Oct 21;12(1):17667. doi: 10.1038/s41598-022-22211-2. Sci Rep. 2022. PMID: 36271283 Free PMC article.
-
Potential biomarkers for predicting the overall survival outcome of kidney renal papillary cell carcinoma: an analysis of ferroptosis-related LNCRNAs.BMC Urol. 2022 Sep 14;22(1):152. doi: 10.1186/s12894-022-01037-0. BMC Urol. 2022. PMID: 36104680 Free PMC article.
-
Overview of the interplay between m6A methylation modification and non-coding RNA and their impact on tumor cells.Transl Cancer Res. 2024 Jun 30;13(6):3106-3125. doi: 10.21037/tcr-23-2401. Epub 2024 Jun 27. Transl Cancer Res. 2024. PMID: 38988908 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

