Coexistence of tet(X4), mcr-1, and blaNDM-5 in ST6775 Escherichia coli Isolates of Animal Origin in China
- PMID: 35311537
- PMCID: PMC9045152
- DOI: 10.1128/spectrum.00196-22
Coexistence of tet(X4), mcr-1, and blaNDM-5 in ST6775 Escherichia coli Isolates of Animal Origin in China
Abstract
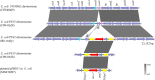
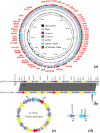
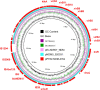
Emergence of pathogens harboring multiple resistance genes incurs great concerns. Cooccurrence of mobile resistance genes conferring resistance to tigecycline, colistin, and carbapenems in Escherichia coli has not been investigated. This study aimed to characterize three E. coli isolates coharboring tet(X4), mcr-1, and blaNDM-5. Isolates coharboring tet(X4), mcr-1, and blaNDM-5 were identified and characterized by PCR, Sanger sequencing, antimicrobial susceptibility testing, conjugation assays, Illumina sequencing, nanopore sequencing, and bioinformatic analysis. Three E. coli isolates carrying tet(X4), mcr-1, and blaNDM-5 were identified from pigeons in China. They were resistant to almost all antimicrobials except enrofloxacin. tet(X4) and blaNDM-5 could be conjugated into E. coli C600, but mcr-1 was nontransferable in three isolates. Three isolates belonged to sequence type 6775 (ST6775), and clonal dissemination of isolates carrying tet(X4), mcr-1, and blaNDM-5 existed in the pigeon farm. Genetic analysis revealed that mcr-1 mediated by the Tn6330 was located on the chromosome, tet(X4) was located on the IncFII plasmid, and blaNDM-5 was located on the IncX3 plasmid. We first characterized the E. coli isolates carrying tet(X4), mcr-1, and blaNDM-5 simultaneously. Relevant measures should be taken to decrease the prevalence of pathogens carrying tet(X4), mcr-1, and blaNDM-5. IMPORTANCE Tigecycline and colistin are regarded as vital antimicrobials to treat multidrug-resistant (MDR) bacterial infections, such as that caused by carbapenemase-producing Enterobacteriaceae (CPE). Cooccurrence of mobile resistance genes conferring resistance to last-resort antimicrobials in E. coli remains unknown. Here, we characterized E. coli strains coharboring tet(X4), mcr-1, and blaNDM-5 phenotypically and genetically. Resistance genes tet(X4), mcr-1, and blaNDM-5 were located on transposons or plasmids that were mobile genetic elements related to the capture, accumulation, and dissemination of such important resistance genes. The emergence of E. coli isolates carrying tet(X4), mcr-1, and blaNDM-5 highlights the importance of monitoring the coexistence of novel mobile resistance genes in different settings with a One Health approach. Risk of transmission of such MDR pathogens from animals to humans should be evaluated comprehensively.
Keywords: Escherichia coli; ST6775; blaNDM-5; coexistence; mcr-1; tet(X4).
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Emerging Opportunity and Destiny of mcr-1- and tet(X4)-Coharboring Plasmids in Escherichia coli.Microbiol Spectr. 2021 Dec 22;9(3):e0152021. doi: 10.1128/Spectrum.01520-21. Epub 2021 Dec 8. Microbiol Spectr. 2021. PMID: 34878308 Free PMC article.
-
Molecular epidemiology and population genomics of tet(X4), blaNDM or mcr-1 positive Escherichia coli from migratory birds in southeast coast of China.Ecotoxicol Environ Saf. 2022 Oct 1;244:114032. doi: 10.1016/j.ecoenv.2022.114032. Epub 2022 Sep 7. Ecotoxicol Environ Saf. 2022. PMID: 36084501
-
Emergence of plasmid-mediated fosfomycin resistance among Escherichia coli harboring fosA4, tet(X4), and mcr-1 genes in wild birds.mSystems. 2025 Apr 22;10(4):e0167324. doi: 10.1128/msystems.01673-24. Epub 2025 Mar 13. mSystems. 2025. PMID: 40079598 Free PMC article.
-
Prevalence and genomic characterization of clinical Escherichia coli strains that harbor the plasmid-borne tet(X4) gene in China.Microbiol Res. 2024 Aug;285:127730. doi: 10.1016/j.micres.2024.127730. Epub 2024 Apr 16. Microbiol Res. 2024. PMID: 38805981 Review.
-
Antimicrobial Resistance in Escherichia coli.Microbiol Spectr. 2018 Jul;6(4):10.1128/microbiolspec.arba-0026-2017. doi: 10.1128/microbiolspec.ARBA-0026-2017. Microbiol Spectr. 2018. PMID: 30003866 Free PMC article. Review.
Cited by
-
Mobile Colistin Resistance (mcr) Gene-Containing Organisms in Poultry Sector in Low- and Middle-Income Countries: Epidemiology, Characteristics, and One Health Control Strategies.Antibiotics (Basel). 2023 Jun 28;12(7):1117. doi: 10.3390/antibiotics12071117. Antibiotics (Basel). 2023. PMID: 37508213 Free PMC article. Review.
-
Whole genome analysis reveals the distribution and diversity of plasmid reservoirs of NDM and MCR in commercial chicken farms in China.Microbiol Spectr. 2025 Jul;13(7):e0290024. doi: 10.1128/spectrum.02900-24. Epub 2025 Jun 9. Microbiol Spectr. 2025. PMID: 40488461 Free PMC article.
-
Bacteria carrying mobile colistin resistance genes and their control measures, an updated review.Arch Microbiol. 2024 Nov 8;206(12):462. doi: 10.1007/s00203-024-04188-w. Arch Microbiol. 2024. PMID: 39516398 Review.
-
Global prevalence, characteristics, and future prospects of IncX3 plasmids: A review.Front Microbiol. 2022 Sep 6;13:979558. doi: 10.3389/fmicb.2022.979558. eCollection 2022. Front Microbiol. 2022. PMID: 36147856 Free PMC article. Review.
-
Mobile Tigecycline Resistance: An Emerging Health Catastrophe Requiring Urgent One Health Global Intervention.Front Microbiol. 2022 Aug 1;13:808744. doi: 10.3389/fmicb.2022.808744. eCollection 2022. Front Microbiol. 2022. PMID: 35979498 Free PMC article. Review.
References
-
- Paul M, Daikos GL, Durante-Mangoni E, Yahav D, Carmeli Y, Benattar YD, Skiada A, Andini R, Eliakim-Raz N, Nutman A, Zusman O, Antoniadou A, Pafundi PC, Adler A, Dickstein Y, Pavleas I, Zampino R, Daitch V, Bitterman R, Zayyad H, Koppel F, Levi I, Babich T, Friberg LE, Mouton JW, Theuretzbacher U, Leibovici L. 2018. Colistin alone versus colistin plus meropenem for treatment of severe infections caused by carbapenem-resistant Gram-negative bacteria: an open-label, randomised controlled trial. Lancet Infect Dis 18:391–400. doi:10.1016/S1473-3099(18)30099-9. - DOI - PubMed
-
- Seifert H, Blondeau J, Dowzicky MJ. 2018. In vitro activity of tigecycline and comparators (2014–2016) among key WHO 'priority pathogens' and longitudinal assessment (2004–2016) of antimicrobial resistance: a report from the T.E.S.T. study. Int J Antimicrob Agents 52:474–484. doi:10.1016/j.ijantimicag.2018.07.003. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

