Exploring the evolution of biochemical models at the network level
- PMID: 35312734
- PMCID: PMC8936491
- DOI: 10.1371/journal.pone.0265735
Exploring the evolution of biochemical models at the network level
Abstract
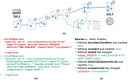
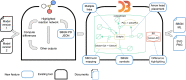
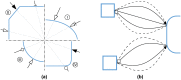
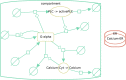
The evolution of biochemical models is difficult to track. At present, it is not possible to inspect the differences between model versions at the network level. Biochemical models are often constructed in a distributed, non-linear process: collaborators create model versions on different branches from novel information, model extensions, during curation and adaption. To discuss and align the versions, it is helpful to abstract the changes to the network level. The differences between two model versions can be detected by the software tool BiVeS. However, it cannot show the structural changes resulting from the differences. Here, we present a method to visualise the differences between model versions effectively. We developed a JSON schema to communicate the differences at the network level and extended BiVeS accordingly. Additionally, we developed DiVil, a web-based tool to represent the model and the differences as a standardised network using D3. It combines an automatic layout with an interactive user interface to improve the visualisation and to inspect the model. The network can be exported in standardised formats as images or markup language. Our method communicates the structural differences between model versions. It facilitates the discussion of changes and thus supports the collaborative and non-linear nature of model development. Availability and implementation: DiVil prototype: https://divil.bio.informatik.uni-rostock.de, Code on GitHub: https://github.com/Gebbi8/DiVil, licensed under Apache License 2.0. Contact: url="tom.gebhardt@uni-rostock.de.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures






Similar articles
-
An algorithm to detect and communicate the differences in computational models describing biological systems.Bioinformatics. 2016 Feb 15;32(4):563-70. doi: 10.1093/bioinformatics/btv484. Epub 2015 Oct 21. Bioinformatics. 2016. PMID: 26490504 Free PMC article.
-
Evolution of computational models in BioModels Database and the Physiome Model Repository.BMC Syst Biol. 2018 Apr 12;12(1):53. doi: 10.1186/s12918-018-0553-2. BMC Syst Biol. 2018. PMID: 29650016 Free PMC article.
-
Development of In-Browser Simulators for Medical Education: Introduction of a Novel Software Toolchain.J Med Internet Res. 2019 Jul 3;21(7):e14160. doi: 10.2196/14160. J Med Internet Res. 2019. PMID: 31271154 Free PMC article.
-
cd2sbgnml: bidirectional conversion between CellDesigner and SBGN formats.Bioinformatics. 2020 Apr 15;36(8):2620-2622. doi: 10.1093/bioinformatics/btz969. Bioinformatics. 2020. PMID: 31904823
-
Reproducible and flexible simulation experiments with ML-Rules and SESSL.Bioinformatics. 2018 Apr 15;34(8):1424-1427. doi: 10.1093/bioinformatics/btx741. Epub 2017 Nov 23. Bioinformatics. 2018. PMID: 29186288
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

