Reductive TCA cycle catalyzed by wild-type IDH2 promotes acute myeloid leukemia and is a metabolic vulnerability for potential targeted therapy
- PMID: 35313945
- PMCID: PMC8935709
- DOI: 10.1186/s13045-022-01245-z
Reductive TCA cycle catalyzed by wild-type IDH2 promotes acute myeloid leukemia and is a metabolic vulnerability for potential targeted therapy
Abstract
Background: Isocitrate dehydrogenase-2 (IDH2) is a mitochondrial enzyme that catalyzes the metabolic conversion between isocitrate and alpha-ketoglutarate (α-KG) in the TCA cycle. IDH2 mutation is an oncogenic event in acute myeloid leukemia (AML) due to the generation of 2-hydroxyglutarate. However, the role of wild-type IDH2 in AML remains unknown, despite patients with it suffer worse clinical outcome than those harboring mutant type.
Methods: IDH2 expression in AML cell lines and patient samples was evaluated by RT-qPCR, western blotting and database analyses. The role of wild-type IDH2 in AML cell survival and proliferation was tested using genetic knockdown and pharmacological inhibition in AML cells and animal models. LC-MS, GC-MS, isotope metabolic tracing, and molecular analyses were performed to reveal the underlying mechanisms.
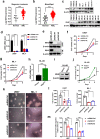
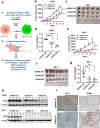
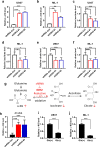
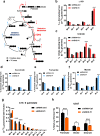
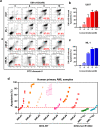
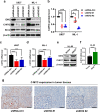
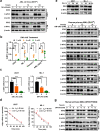
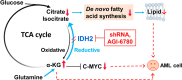
Results: We found that wild-type IDH2 was overexpressed in AML and played a major role in promoting leukemia cell survival and proliferation in vitro and in vivo. Metabolomic analyses revealed an active IDH2-mediated reductive TCA cycle that promoted the conversion of α-KG to isocitrate/citrate to facilitate glutamine utilization for lipid synthesis in AML cells. Suppression of wild-type IDH2 by shRNA resulted in elevated α-KG and decreased isocitrate/citrate, leading to reduced lipid synthesis, a significant decrease in c-Myc downregulated by α-KG, and an inhibition of AML viability and proliferation. Importantly, pharmacological inhibition of IDH2 showed significant therapeutic effect in mice inoculated with AML cells with wt-IDH2 and induced a downregulation of C-MYC in vivo.
Conclusions: Wt-IDH2 is an essential molecule for AML cell survival and proliferation by promoting conversion of α-KG to isocitrate for lipid synthesis and by upregulating c-Myc expression and could be a potential therapeutic target in AML.
Keywords: Acute myeloid leukemia; Alpha-ketoglutarate; Lipid synthesis; TCA cycle; Wild-type IDH2; c-Myc.
© 2022. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Isocitrate dehydrogenase 2 mutation promotes cytarabine resistance in acute myeloid leukemia by Warburg effect.Hematol Oncol. 2024 Nov;42(6):e3316. doi: 10.1002/hon.3316. Hematol Oncol. 2024. PMID: 39526588 Free PMC article.
-
All-Trans Retinoic Acid Synergizes with Enasidenib to Induce Differentiation of IDH2-Mutant Acute Myeloid Leukemia Cells.Yonsei Med J. 2020 Sep;61(9):762-773. doi: 10.3349/ymj.2020.61.9.762. Yonsei Med J. 2020. PMID: 32882760 Free PMC article.
-
IDH2 inhibition in AML: Finally progress?Best Pract Res Clin Haematol. 2015 Jun-Sep;28(2-3):112-5. doi: 10.1016/j.beha.2015.10.016. Epub 2015 Oct 19. Best Pract Res Clin Haematol. 2015. PMID: 26590767 Review.
-
Wild-type IDH2 is a therapeutic target for triple-negative breast cancer.Nat Commun. 2024 Apr 24;15(1):3445. doi: 10.1038/s41467-024-47536-6. Nat Commun. 2024. PMID: 38658533 Free PMC article.
-
Targeting the IDH2 Pathway in Acute Myeloid Leukemia.Clin Cancer Res. 2018 Oct 15;24(20):4931-4936. doi: 10.1158/1078-0432.CCR-18-0536. Epub 2018 May 16. Clin Cancer Res. 2018. PMID: 29769206 Review.
Cited by
-
Targeting Mitochondrial IDH2 Enhances Antitumor Activity of Cisplatin in Lung Cancer via ROS-Mediated Mechanism.Biomedicines. 2023 Feb 7;11(2):475. doi: 10.3390/biomedicines11020475. Biomedicines. 2023. PMID: 36831011 Free PMC article.
-
Inhibition of IDH3α Enhanced the Efficacy of Chemoimmunotherapy by Regulating Acidic Tumor Microenvironments.Cancers (Basel). 2023 Mar 16;15(6):1802. doi: 10.3390/cancers15061802. Cancers (Basel). 2023. PMID: 36980689 Free PMC article.
-
Endometriotic lesions exhibit distinct metabolic signature compared to paired eutopic endometrium at the single-cell level.Commun Biol. 2024 Aug 21;7(1):1026. doi: 10.1038/s42003-024-06713-5. Commun Biol. 2024. PMID: 39169201 Free PMC article.
-
Wild-Type Isocitrate Dehydrogenase-Dependent Oxidative Decarboxylation and Reductive Carboxylation in Cancer and Their Clinical Significance.Cancers (Basel). 2022 Nov 24;14(23):5779. doi: 10.3390/cancers14235779. Cancers (Basel). 2022. PMID: 36497259 Free PMC article. Review.
-
Nucleolin recognizing silica nanoparticles inhibit cell proliferation by activating the Bax/Bcl-2/caspase-3 signalling pathway to induce apoptosis in liver cancer.Front Pharmacol. 2023 Feb 9;14:1117052. doi: 10.3389/fphar.2023.1117052. eCollection 2023. Front Pharmacol. 2023. PMID: 36843953 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

